Ann Dermatol.
2009 Nov;21(4):352-357. 10.5021/ad.2009.21.4.352.
Comparison of Nested PCR and RFLP for Identification and Classification of Malassezia Yeasts from Healthy Human Skin
- Affiliations
-
- 1Department of Dermatology, Konkuk University School of Medicine, Seoul, Korea. 20050078@kuh.ac.kr
- KMID: 2156496
- DOI: http://doi.org/10.5021/ad.2009.21.4.352
Abstract
- BACKGROUND
Malassezia yeasts are normal flora of the skin found in 75~98% of healthy subjects. The accurate identification of the Malassezia species is important for determining the pathogenesis of the Malassezia yeasts with regard to various skin diseases such as Malassezia folliculitis, seborrheic dermatitis, and atopic dermatitis.
OBJECTIVE
This research was conducted to determine a more accurate and rapid molecular test for the identification and classification of Malassezia yeasts.
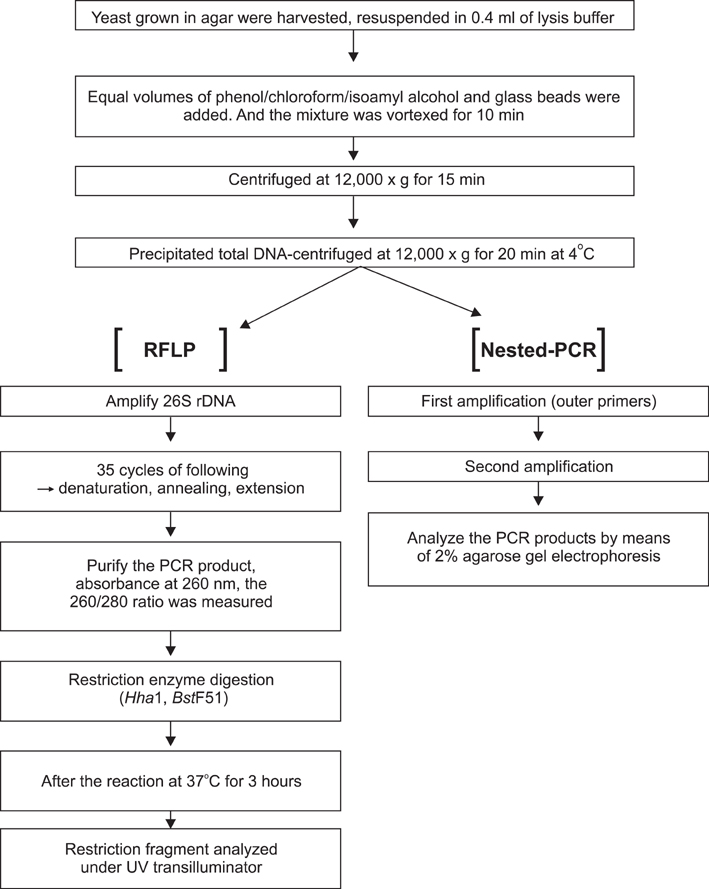
METHODS
We compared the accuracy and efficacy of restriction fragment length polymorphism (RFLP) and the nested polymerase chain reaction (PCR) for the identification of Malassezia yeasts.
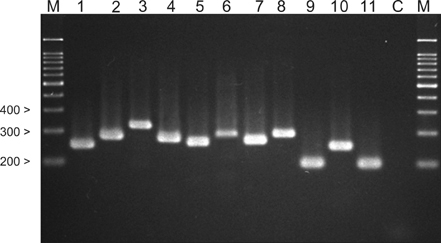
RESULTS
Although both methods demonstrated rapid and reliable results with regard to identification, the nested PCR method was faster. However, 7 different Malassezia species (1.2%) were identified by the nested PCR compared to the RFLP method.
CONCLUSION
Our results show that RFLP method was relatively more accurate and reliable for the detection of various Malassezia species compared to the nested PCR. But, in the aspect of simplicity and time saving, the latter method has its own advantages. In addition, the 26S rDNA, which was targeted in this study, contains highly conserved base sequences and enough sequence variation for inter-species identification of Malassezia yeasts.
MeSH Terms
Figure
Cited by 3 articles
-
Molecular Analysis of Malassezia Microflora on the Skin of the Patients with Atopic Dermatitis
Seon Mi Yim, Ji Young Kim, Jong Hyun Ko, Yang Won Lee, Yong Beom Choe, Kyu Joong Ahn
Ann Dermatol. 2010;22(1):41-47. doi: 10.5021/ad.2010.22.1.41.Molecular Biological Identification of Malassezia Yeasts Using Pyrosequencing
Ji Young Kim, Hyung Jin Hahn, Yong Beom Choe, Yang Won Lee, Kyu Joong Ahn, Kee Chan Moon
Ann Dermatol. 2013;25(1):73-79. doi: 10.5021/ad.2013.25.1.73.Progress in Malassezia Research in Korea
Soo Young Kim, Yang Won Lee, Yong Beom Choe, Kyu Joong Ahn
Ann Dermatol. 2015;27(6):647-657. doi: 10.5021/ad.2015.27.6.647.
Reference
-
1. Gupta AK, Batra R, Bluhm R, Boekhout T, Dawson TL Jr. Skin diseases associated with Malassezia species. J Am Acad Dermatol. 2004. 51:785–798.2. Gueho E, Midgley G, Guillot J. The genus Malassezia with description of four new species. Antonie Van Leeuwenhoek. 1996. 69:337–355.3. Sugita T, Kodama M, Saito M, Ito T, Kato Y, Tsuboi R, et al. Sequence diversity of the intergenic spacer region of the rRNA gene of Malassezia globosa colonizing the skin of patients with atopic dermatitis and healthy individuals. J Clin Microbiol. 2003. 41:3022–3027.
Article4. Sugita T, Takashima M, Shinoda T, Suto H, Unno T, Tsuboi R, et al. New yeast species, Malassezia dermatis, isolated from patients with atopic dermatitis. J Clin Microbiol. 2002. 40:1363–1367.
Article5. Sugita T, Takashima M, Kodama M, Tsuboi R, Nishikawa A. Description of a new yeast species, Malassezia japonica, and its detection in patients with atopic dermatitis and healthy subjects. J Clin Microbiol. 2003. 41:4695–4699.
Article6. Hirai A, Kano R, Makimura K, Duarte ER, Hamdan JS, Lachance MA, et al. Malassezia nana sp. nov., a novel lipid-dependent yeast species isolated from animals. Int J Syst Evol Microbiol. 2004. 54:623–627.
Article7. Sugita T, Tajima M, Takashima M, Amaya M, Saito M, Tsuboi R, et al. A new yeast, Malassezia yamatoensis, isolated from a patient with seborrheic dermatitis, and its distribution in patients and healthy subjects. Microbiol Immunol. 2004. 48:579–583.
Article8. Lee YW, Yim SM, Lim SH, Choe YB, Ahn KJ. Quantitative investigation on the distribution of Malassezia species on healthy human skin in Korea. Mycoses. 2006. 49:405–410.
Article9. Guillot J, Gueho E, Lesourd M, Midgley G, Chevrier G, Dupont B. Identification of Malassezia species. A practical approach. J Mycol Med. 1996. 6:103–110.10. Rodriguez-Valero S, Mesa LM, Gonzalez-Moran E, Delmonte ML, Robertiz S, Valero A. Phenotypic characterization of species of Malassezia in healthy skin of an university student population. Invest Clin. 2005. 46:329–335.11. Bernier V, Weill FX, Hirigoyen V, Elleau C, Feyler A, Labreze C, et al. Skin colonization by Malassezia species in neonates: a prospective study and relationship with neonatal cephalic pustulosis. Arch Dermatol. 2002. 138:215–218.12. Morishita N, Sei Y, Sugita T. Molecular analysis of Malassezia microflora from patients with pityriasis versicolor. Mycopathologia. 2006. 161:61–65.
Article13. Sugita T, Tajima M, Tsubuku H, Tsuboi R, Nishikawa A. Quantitative analysis of cutaneous Malassezia in atopic dermatitis patients using real-time PCR. Microbiol Immunol. 2006. 50:549–552.
Article14. Senczek D, Siesenop U, Bohm KH. Characterization of Malassezia species by means of phenotypic characteristics and detection of electrophoretic karyotypes by pulsed-field gel electrophoresis (PFGE). Mycoses. 1999. 42:409–414.15. Gupta AK, Boekhout T, Theelen B, Summerbell R, Batra R. Identification and typing of Malassezia species by amplified fragment length polymorphism and sequence analyses of the internal transcribed spacer and large-subunit regions of ribosomal DNA. J Clin Microbiol. 2004. 42:4253–4260.
Article16. Theelen B, Silvestri M, Gueho E, van Belkum A, Boekhout T. Identification and typing of Malassezia yeasts using amplified fragment length polymorphism (AFLP), random amplified polymorphic DNA (RAPD) and denaturing gradient gel electrophoresis (DGGE). FEMS Yeast Res. 2001. 1:79–86.
Article17. Gandra RF, Simao RC, Matsumoto FE, da Silva BC, Ruiz LS, da Silva EG, et al. Genotyping by RAPD-PCR analyses of Malassezia furfur strains from pityriasis versicolor and seborrhoeic dermatitis patients. Mycopathologia. 2006. 162:273–280.
Article18. Gaitanis G, Velegraki A, Alexopoulos EC, Chasapi V, Tsigonia A, Katsambas A. Distribution of Malassezia species in pityriasis versicolor and seborrhoeic dermatitis in Greece. Typing of the major pityriasis versicolor isolate M. globosa. Br J Dermatol. 2006. 154:854–859.
Article19. Gemmer CM, DeAngelis YM, Theelen B, Boekhout T, Dawson TL Jr. Fast, noninvasive method for molecular detection and differentiation of Malassezia yeast species on human skin and application of the method to dandruff microbiology. J Clin Microbiol. 2002. 40:3350–3357.
Article20. Guillot J, Deville M, Berthelemy M, Provost F, Gueho E. A single PCR-restriction endonuclease analysis for rapid identification of Malassezia species. Lett Appl Microbiol. 2000. 31:400–403.
Article21. Makimura K, Tamura Y, Kudo M, Uchida K, Saito H, Yamaguchi H. Species identification and strain typing of Malassezia species stock strains and clinical isolates based on the DNA sequences of nuclear ribosomal internal transcribed spacer 1 regions. J Med Microbiol. 2000. 49:29–35.
Article22. Gaitanis G, Velegraki A, Frangoulis E, Mitroussia A, Tsigonia A, Tzimogianni A, et al. Identification of Malassezia species from patient skin scales by PCR-RFLP. Clin Microbiol Infect. 2002. 8:162–173.
Article23. Mirhendi H, Makimura K, Zomorodian K, Yamada T, Sugita T, Yamaguchi H. A simple PCR-RFLP method for identification and differentiation of 11 Malassezia species. J Microbiol Methods. 2005. 61:281–284.
Article24. Lee YW, Lim SH, Ahn KJ. The application of 26S rDNA PCR-RFLP in the identification and classification of Malassezia yeast. Korean J Med Mycol. 2006. 11:141–153.25. Casagrande BF, Fluckiger S, Linder MT, Johansson C, Scheynius A, Crameri R, et al. Sensitization to the yeast Malassezia sympodialis is specific for extrinsic and intrinsic atopic eczema. J Invest Dermatol. 2006. 126:2414–2421.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Progress in Malassezia Research in Korea
- Epidemiologic Study of Malassezia Yeasts in Patients with Malassezia Folliculitis by 26S rDNA PCR-RFLP Analysis
- The Application of 26S rDNA PCR-RFLP in the Identification and Classification of Malassezia Yeast
- Epidemiologic Study of Malassezia Yeasts in Acne Patients by Analysis of 26S rDNA PCR-RFLP
- Epidemiologic Study of Malassezia Yeasts in Seborrheic Dermatitis Patients by the Analysis of 26S rDNA PCR-RFLP