Yonsei Med J.
2013 Jan;54(1):177-182. 10.3349/ymj.2013.54.1.177.
Molecular and Epidemiological Characterization of Carbapenem-Resistant Acinetobacter baumannii in Non-Tertiary Korean Hospitals
- Affiliations
-
- 1Division of Antimicrobial Resistance, Center for Infectious Diseases, Korea National Institute of Health, Cheongwon, Korea. gtchung@nih.go.kr
- KMID: 1776935
- DOI: http://doi.org/10.3349/ymj.2013.54.1.177
Abstract
- PURPOSE
The increasing prevalence and global spread of carbapenem-resistant Acinetobacter baumannii (A. baumannii) has become a serious problem. The aim of this study was to investigate molecular and epidemiological characteristics of carbapenem-resistant A. baumannii isolates collected from Korean non-tertiary hospitals.
MATERIALS AND METHODS
Thirty six non-duplicated carbapenem-resistant A. baumannii isolates were collected from 17 non-tertiary hospitals in Korea between 2004 and 2006. Isolates were typed by multilocus sequence typing and repetitive-sequence-based PCR (rep-PCR). Detection of genes encoding OXA carbapenemase and their relationship with ISAba1 was performed by PCR.
RESULTS
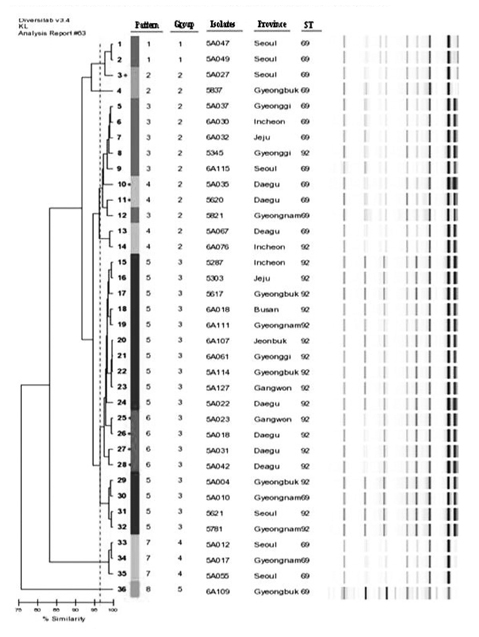
Two clones were prevalent among 36 isolates: ST69 (17 isolates, 47.2%) and ST92 (19 isolates, 52.8%). Rep-PCR patterns were diverse and revealed that all isolates were clustered into eight band patterns. The ISAba1-activated blaOXA-23-like and ISAba1-activated blaOXA-51-like genes were prevalent among the carbapenem-resistant A. baumannii isolates.
CONCLUSION
The class D beta-lactamase genes of A. baumannii were distributed nationwide in non-tertiary Korean hospitals.
MeSH Terms
-
Acinetobacter Infections/epidemiology/*microbiology
Acinetobacter baumannii/classification/*genetics
Anti-Bacterial Agents/therapeutic use
Bacterial Typing Techniques
Carbapenems/*therapeutic use
DNA, Bacterial/analysis
*Drug Resistance, Bacterial
Hospitals
Humans
Microbial Sensitivity Tests
Molecular Epidemiology
Multilocus Sequence Typing
Polymerase Chain Reaction
Prevalence
Republic of Korea
beta-Lactamases/genetics
Anti-Bacterial Agents
Carbapenems
DNA, Bacterial
beta-Lactamases
Figure
Reference
-
1. Munoz-Price LS, Weinstein RA. Acinetobacter infection. N Engl J Med. 2008. 358:1271–1281.
Article2. Poirel L, Lebessi E, Héritier C, Patsoura A, Foustoukou M, Nordmann P. Nosocomial spread of OXA-58-positive carbapenem-resistant Acinetobacter baumannii isolates in a paediatric hospital in Greece. Clin Microbiol Infect. 2006. 12:1138–1141.
Article3. Garnacho-Montero J, Amaya-Villar R. Multiresistant Acinetobacter baumannii infections: epidemiology and management. Curr Opin Infect Dis. 2010. 23:332–339.
Article4. Kwon KT, Oh WS, Song JH, Chang HH, Jung SI, Kim SW, et al. Impact of imipenem resistance on mortality in patients with Acinetobacter bacteraemia. J Antimicrob Chemother. 2007. 59:525–530.
Article5. Lee JH, Choi CH, Kang HY, Lee JY, Kim J, Lee YC, et al. Differences in phenotypic and genotypic traits against antimicrobial agents between Acinetobacter baumannii and Acinetobacter genomic species 13TU. J Antimicrob Chemother. 2007. 59:633–639.
Article6. Walther-Rasmussen J, Høiby N. OXA-type carbapenemases. J Antimicrob Chemother. 2006. 57:373–383.
Article7. Park YK, Lee GH, Baek JY, Chung DR, Peck KR, Song JH, et al. A single clone of Acinetobacter baumannii, ST22, is responsible for high antimicrobial resistance rates of Acinetobacter spp. isolates that cause bacteremia and urinary tract infections in Korea. Microb Drug Resist. 2010. 16:143–149.
Article8. Park YK, Choi JY, Jung SI, Park KH, Lee H, Jung DS, et al. Two distinct clones of carbapenem-resistant Acinetobacter baumannii isolates from Korean hospitals. Diagn Microbiol Infect Dis. 2009. 64:389–395.
Article9. La Scola B, Gundi VA, Khamis A, Raoult D. Sequencing of the rpoB gene and flanking spacers for molecular identification of Acinetobacter species. J Clin Microbiol. 2006. 44:827–832.
Article10. Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing: Twentieth Informational Supplement M100-S20. 2010. Wayne, PA, USA: CLSI.11. Bartual SG, Seifert H, Hippler C, Luzon MA, Wisplinghoff H, Rodríguez-Valera F. Development of a multilocus sequence typing scheme for characterization of clinical isolates of Acinetobacter baumannii. J Clin Microbiol. 2005. 43:4382–4390.
Article12. Feil EJ, Li BC, Aanensen DM, Hanage WP, Spratt BG. eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J Bacteriol. 2004. 186:1518–1530.
Article13. Woodford N, Ellington MJ, Coelho JM, Turton JF, Ward ME, Brown S, et al. Multiplex PCR for genes encoding prevalent OXA carbapenemases in Acinetobacter spp. Int J Antimicrob Agents. 2006. 27:351–353.
Article14. Manikal VM, Landman D, Saurina G, Oydna E, Lal H, Quale J. Endemic carbapenem-resistant Acinetobacter species in Brooklyn, New York: citywide prevalence, interinstitutional spread, and relation to antibiotic usage. Clin Infect Dis. 2000. 31:101–106.
Article15. Ko KS, Suh JY, Kwon KT, Jung SI, Park KH, Kang CI, et al. High rates of resistance to colistin and polymyxin B in subgroups of Acinetobacter baumannii isolates from Korea. J Antimicrob Chemother. 2007. 60:1163–1167.
Article16. Lee Y, Lee J, Jeong SH, Lee J, Bae IK, Lee K. Carbapenem-non-susceptible Acinetobacter baumannii of sequence type 92 or its single-locus variants with a G428T substitution in zone 2 of the rpoB gene. J Antimicrob Chemother. 2011. 66:66–72.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Carbapenem-Resistant Acinetobacter baumannii
- ISAba15 Inserted into Outer Membrane Protein Gene carO in Acinetobacter baumannii
- Epidemiological characteristics of carbapenemresistant Enterobacteriaceae and carbapenem-resistant Acinetobacter baumannii in a tertiary referral hospital in Korea
- Update on the Epidemiology, Treatment, and Outcomes of Carbapenem-resistant Acinetobacter infections
- Molecular typing of Acinetobacter baumannii strains by randomly amplified polymorphic DNA (RAPD) analysis