Cancer Res Treat.
2015 Oct;47(4):796-803. 10.4143/crt.2014.106.
Can Serum be Used for Analyzing the KRAS Mutation Status in Patients with Advanced Colorectal Cancer?
- Affiliations
-
- 1Division of Hematology-Oncology, Department of Medicine, Korea University College of Medicine, Seoul, Korea. yhk0215@korea.ac.kr
- 2Division of Hematology-Oncology, Department of Medicine, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea.
- 3Genomic Research Center for Lung and Breast/Ovarian Cancers, Korea University Anam Hospital, Seoul, Korea.
- KMID: 2403399
- DOI: http://doi.org/10.4143/crt.2014.106
Abstract
- PURPOSE
KRAS mutations have been used widely as prognostic or predictive marker in patients with advanced colorectal cancer (CRC). However, it may be difficult to obtain a tumor tissue for analyzing the status of KRAS mutation in large proportion of patients with advanced disease.
MATERIALS AND METHODS
We obtained pairs of tumor and serum samples from 65 patients with advanced CRC, between March 2008 and July 2011. KRAS mutation status from the tumor samples was analyzed by genomic polymerase chain reaction and direct sequence, and KRASmutation status from the serum samples was determined by a genomic polymerase chain reaction-restriction fragment length polymorphism assay.
RESULTS
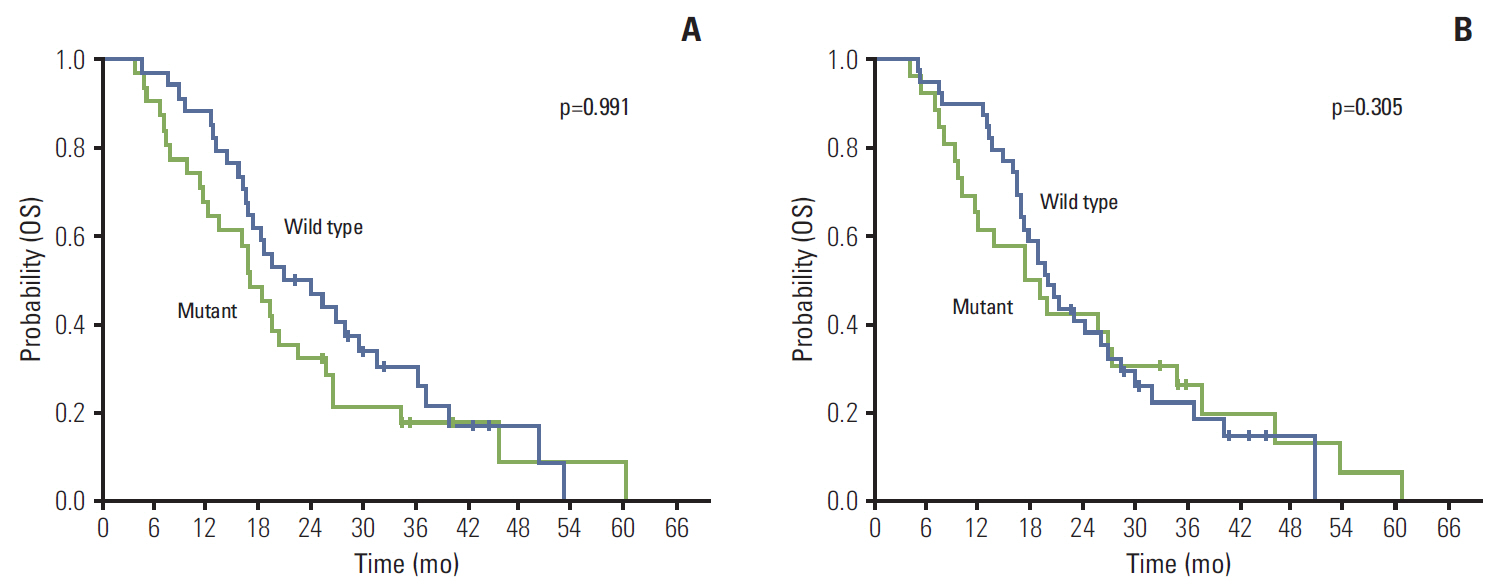
KRAS mutations were detected in the serum samples of 26 patients and in the tumor samples of 31 patients. KRAS mutation status in the serum and tumor samples was consistent in 44 of the 65 pairs (67.7%). There was a significant correlation between the mutations detected in the serum sample and the mutations detected in the matched tumor sample (correlation index, 0.35; p < 0.004). Twenty-two of the 57 patients (38.5%) received anti-epidermal growth factor receptor therapy as any line therapy. There was no significant difference in the overall survival (OS) in accordance to the status of KRASmutations in both the serum and tumor samples (p > 0.05). In a multivariate analysis, liver metastasis and no cytoreductive operation were independent prognostic factors for decreased OS.
CONCLUSION
The serum sample might alternatively be used when it is difficult to obtain tumor tissues for analyzing the status of KRAS mutation in patients with advanced CRC.
MeSH Terms
Figure
Reference
-
References
1. Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005; 55:74–108.
Article2. Jiang Y, Kimchi ET, Staveley-O'Carroll KF, Cheng H, Ajani JA. Assessment of K-ras mutation: a step toward personalized medicine for patients with colorectal cancer. Cancer. 2009; 115:3609–17.3. Andreyev HJ, Norman AR, Cunningham D, Oates J, Dix BR, Iacopetta BJ, et al. Kirsten ras mutations in patients with colorectal cancer: the 'RASCAL II' study. Br J Cancer. 2001; 85:692–6.4. Bazan V, Migliavacca M, Zanna I, Tubiolo C, Grassi N, Latteri MA, et al. Specific codon 13 K-ras mutations are predictive of clinical outcome in colorectal cancer patients, whereas codon 12 K-ras mutations are associated with mucinous histotype. Ann Oncol. 2002; 13:1438–46.
Article5. Schubbert S, Shannon K, Bollag G. Hyperactive Ras in developmental disorders and cancer. Nat Rev Cancer. 2007; 7:295–308.
Article6. Poehlmann A, Kuester D, Meyer F, Lippert H, Roessner A, Schneider-Stock R. K-ras mutation detection in colorectal cancer using the pyrosequencing technique. Pathol Res Pract. 2007; 203:489–97.
Article7. Costa DB, Kobayashi S, Tenen DG, Huberman MS. Pooled analysis of the prospective trials of gefitinib monotherapy for EGFR-mutant non-small cell lung cancers. Lung Cancer. 2007; 58:95–103.
Article8. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011; 11:426–37.
Article9. Gormally E, Caboux E, Vineis P, Hainaut P. Circulating free DNA in plasma or serum as biomarker of carcinogenesis: practical aspects and biological significance. Mutat Res. 2007; 635:105–17.
Article10. Kopreski MS, Benko FA, Kwee C, Leitzel KE, Eskander E, Lipton A, et al. Detection of mutant K-ras DNA in plasma or serum of patients with colorectal cancer. Br J Cancer. 1997; 76:1293–9.
Article11. Anker P, Lefort F, Vasioukhin V, Lyautey J, Lederrey C, Chen XQ, et al. K-ras mutations are found in DNA extracted from the plasma of patients with colorectal cancer. Gastroenterology. 1997; 112:1114–20.
Article12. Amado RG, Wolf M, Peeters M, Van Cutsem E, Siena S, Freeman DJ, et al. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol. 2008; 26:1626–34.13. Karapetis CS, Khambata-Ford S, Jonker DJ, O'Callaghan CJ, Tu D, Tebbutt NC, et al. K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N Engl J Med. 2008; 359:1757–65.14. Andreyev HJ, Norman AR, Cunningham D, Oates JR, Clarke PA. Kirsten ras mutations in patients with colorectal cancer: the multicenter "RASCAL" study. J Natl Cancer Inst. 1998; 90:675–84.
Article15. Hurwitz H, Fehrenbacher L, Novotny W, Cartwright T, Hainsworth J, Heim W, et al. Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N Engl J Med. 2004; 350:2335–42.
Article16. Valent A, Penault-Llorca F, Cayre A, Kroemer G. Change in HER2 (ERBB2) gene status after taxane-based chemotherapy for breast cancer: polyploidization can lead to diagnostic pitfalls with potential impact for clinical management. Cancer Genet. 2013; 206:37–41.17. Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013; 368:1199–209.
Article18. Zlobec I, Bihl MP, Schwarb H, Terracciano L, Lugli A. Clinicopathological and protein characterization of BRAF- and K-RAS-mutated colorectal cancer and implications for prognosis. Int J Cancer. 2010; 127:367–80.19. Naguib A, Mitrou PN, Gay LJ, Cooke JC, Luben RN, Ball RY, et al. Dietary, lifestyle and clinicopathological factors associated with BRAF and K-ras mutations arising in distinct subsets of colorectal cancers in the EPIC Norfolk study. BMC Cancer. 2010; 10:99.
Article20. Gonzalez-Aguilera JJ, Oliart S, Azcoita MM, Fernandez-Peralta AM. Simultaneous mutations in K-ras and TP53 are indicative of poor prognosis in sporadic colorectal cancer. Am J Clin Oncol. 2004; 27:39–45.21. Westra JL, Schaapveld M, Hollema H, de Boer JP, Kraak MM, de Jong D, et al. Determination of TP53 mutation is more relevant than microsatellite instability status for the prediction of disease-free survival in adjuvant-treated stage III colon cancer patients. J Clin Oncol. 2005; 23:5635–43.
Article22. Leslie A, Pratt NR, Gillespie K, Sales M, Kernohan NM, Smith G, et al. Mutations of APC, K-ras, and p53 are associated with specific chromosomal aberrations in colorectal adenocarcinomas. Cancer Res. 2003; 63:4656–61.23. Baker SJ, Preisinger AC, Jessup JM, Paraskeva C, Markowitz S, Willson JK, et al. p53 gene mutations occur in combination with 17p allelic deletions as late events in colorectal tumorigenesis. Cancer Res. 1990; 50:7717–22.24. Fearon ER, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990; 61:759–67.
Article25. Klein CA. Parallel progression of primary tumours and metastases. Nat Rev Cancer. 2009; 9:302–12.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Impact of KRAS Mutation Status on Outcomes in Metastatic Colon Cancer Patients without Anti-Epidermal Growth Factor Receptor Therapy
- Dose KRAS Mutation Status Affect on the Effect of VEGF Therapy in Metastatic Colon Cancer Patients?
- Magnetic Resonance-Based Texture Analysis Differentiating KRAS Mutation Status in Rectal Cancer
- Rapid and Sensitive Detection of KRAS Mutation by Peptide Nucleic Acid-based Real-time PCR Clamping: A Comparison with Direct Sequencing between Fresh Tissue and Formalin-fixed and Paraffin Embedded Tissue of Colorectal Cancer
- p16 Hypermethylation and KRAS Mutation Are Independent Predictors of Cetuximab Plus FOLFIRI Chemotherapy in Patients with Metastatic Colorectal Cancer