Intest Res.
2019 Jul;17(3):387-397. 10.5217/ir.2018.00167.
Quantitative histology-based classification system for assessment of the intestinal mucosal histological changes in patients with celiac disease
- Affiliations
-
- 1Department of Pathology, All India Institute of Medical Sciences, New Delhi, India. prasenaiims@gmail.com
- 2Department of Gastroenterology and Human Nutrition, All India Institute of Medical Sciences, New Delhi, India. govindmakharia@gmail.com
- 3Department of Biostatistics, All India Institute of Medical Sciences, New Delhi, India.
- KMID: 2454747
- DOI: http://doi.org/10.5217/ir.2018.00167
Abstract
- BACKGROUND/AIMS
The existing histological classifications for the interpretation of small intestinal biopsies are based on qualitative parameters with high intraobserver and interobserver variations. We have developed and propose a quantitative histological classification system for the assessment of intestinal mucosal biopsies.
METHODS
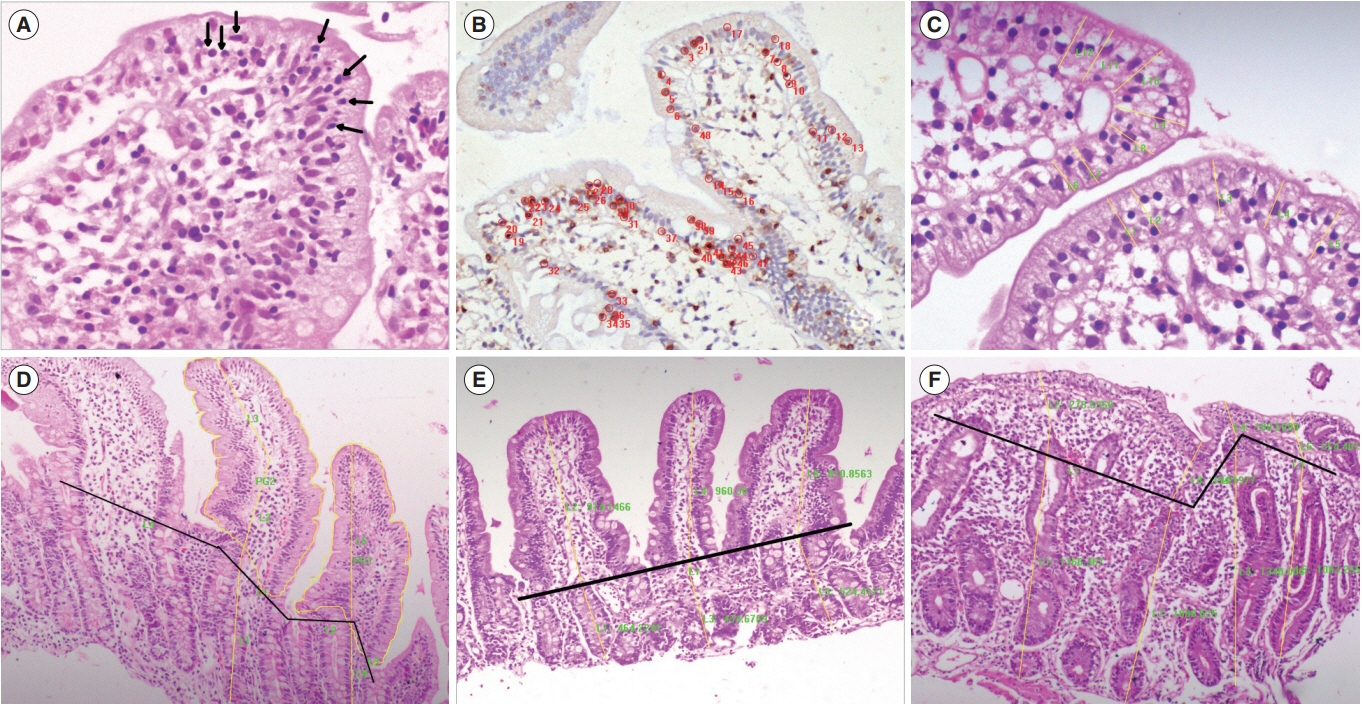
We performed a computer-assisted quantitative histological assessment of digital images of duodenal biopsies from 137 controls and 124 patients with celiac disease (CeD) (derivation cohort). From the receiver-operating curve analysis, followed by multivariate and logistic regression analyses, we identified parameters for differentiating control biopsies from those of the patients with CeD. We repeated the quantitative histological analysis in a validation cohort (105 controls and 120 patients with CeD). On the basis of the results, we propose a quantitative histological classification system. The new classification was compared with the existing histological classifications for interobserver and intraobserver agreements by a group of qualified pathologists.
RESULTS
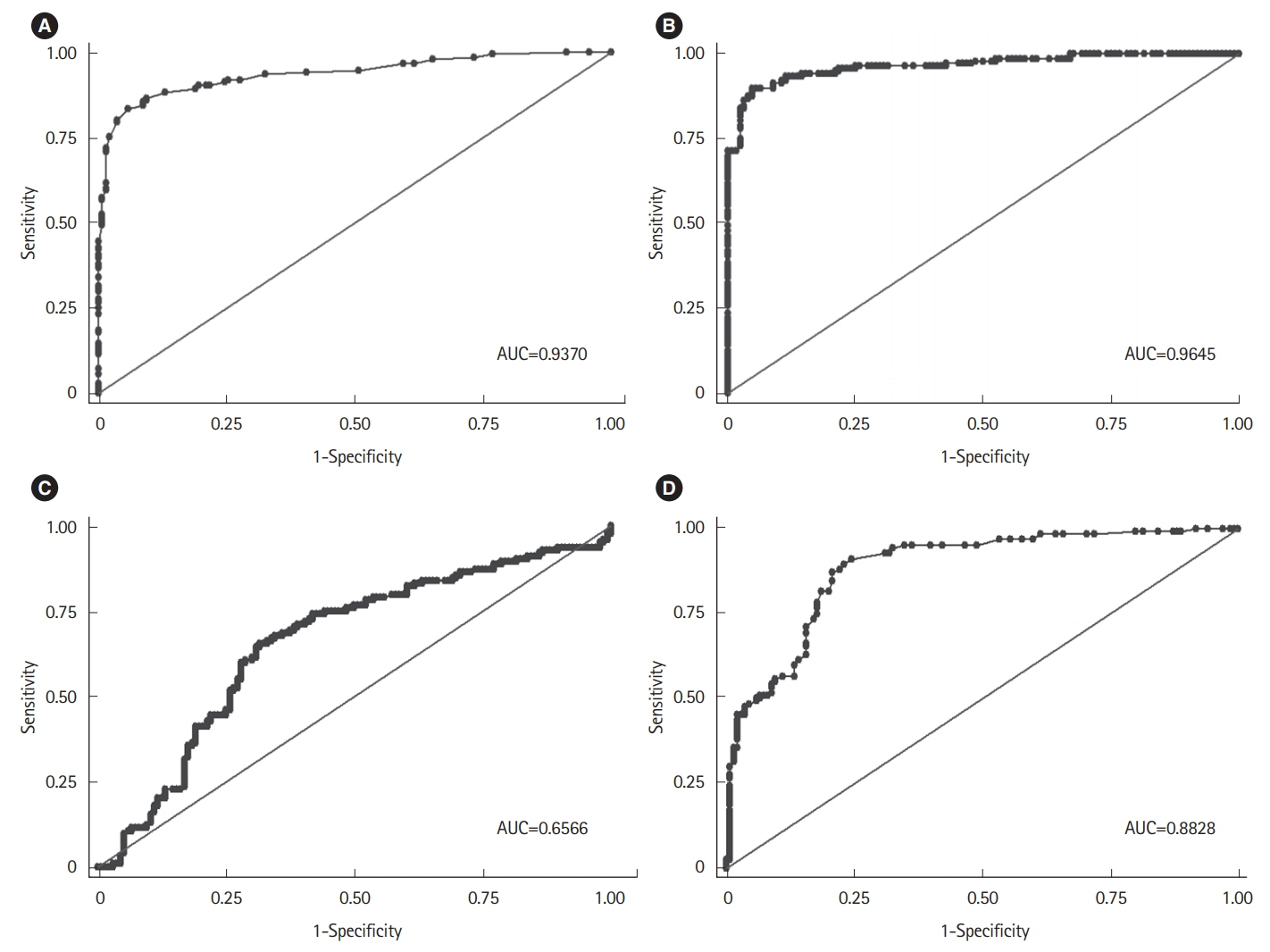
Among the histological parameters, intraepithelial lymphocyte count of ≥25/100 epithelial cells, adjusted villous height fold change of ≤0.7, and crypt depth-to-villous height ratio of ≥0.5 showed good discriminative power between the mucosal biopsies from the patients with CeD and those from the controls, with 90.3% sensitivity, 93.5% specificity, and 96.2% area under the curve. Among the existing histological classifications, our quantitative histological classification showed the highest intraobserver (69.7%-85.03%) and interobserver (24.6%-71.5%) agreements.
CONCLUSIONS
Quantitative assessment increases the reliability of the histological assessment of mucosal biopsies in patients with CeD. Such a classification system may be used for clinical trials in patients with CeD.
MeSH Terms
Figure
Reference
-
1. Oberhuber G, Granditsch G, Vogelsang H. The histopathology of coeliac disease: time for a standardized report scheme for pathologists. Eur J Gastroenterol Hepatol. 1999; 11:1185–1194.2. Marsh MN. Gluten, major histocompatibility complex, and the small intestine. A molecular and immunobiologic approach to the spectrum of gluten sensitivity (‘celiac sprue’). Gastroenterology. 1992; 102:330–354.
Article3. Revised criteria for diagnosis of coeliac disease. Report of Working Group of European Society of Paediatric Gastroenterology and Nutrition. Arch Dis Child. 1990; 65:909–911.4. Madan M, Piplani S, Sharma M, Bhasin TS, Manjari M, Kaur H. Celiac disease: an assessment of subjective variation and diagnostic reproducibility of the various classification systems. Glob J Med Res. 2015; 15(No 1-C):882.5. Corazza GR, Villanacci V, Zambelli C, et al. Comparison of the interobserver reproducibility with different histologic criteria used in celiac disease. Clin Gastroenterol Hepatol. 2007; 5:838–843.
Article6. Ensari A. Gluten-sensitive enteropathy (celiac disease): controversies in diagnosis and classification. Arch Pathol Lab Med. 2010; 134:826–836.
Article7. Cummins AG, Alexander BG, Chung A, et al. Morphometric evaluation of duodenal biopsies in celiac disease. Am J Gastroenterol. 2011; 106:145–150.
Article8. Meinhard EA, Wadbrook DG, Risdon RA. Computer card morphometry of jejunal biopsies in childhood coeliac disease. J Clin Pathol. 1975; 28:85–93.
Article9. Taavela J, Koskinen O, Huhtala H, et al. Validation of morphometric analyses of small-intestinal biopsy readouts in celiac disease. PLoS One. 2013; 8:e76163.
Article10. Kuitunen P, Kosnai I, Savilahti E. Morphometric study of the jejunal mucosa in various childhood enteropathies with special reference to intraepithelial lymphocytes. J Pediatr Gastroenterol Nutr. 1982; 1:525–531.
Article11. Ciclitira PJ, Evans DJ, Fagg NL, Lennox ES, Dowling RH. Clinical testing of gliadin fractions in coeliac patients. Clin Sci (Lond). 1984; 66:357–364.
Article12. Tulloh EA, Baylis JM, Challacombe DN. Automated analysis of morphological change in the duodenal mucosa of children with coeliac disease. Arch Dis Child. 1981; 56:860–863.
Article13. Ghoshal UC, Gwee KA. Post-infectious IBS, tropical sprue and small intestinal bacterial overgrowth: the missing link. Nat Rev Gastroenterol Hepatol. 2017; 14:435–441.
Article14. Husby S, Koletzko S, Korponay-Szabó IR, et al. European Society for Pediatric Gastroenterology, Hepatology, and Nutrition guidelines for the diagnosis of coeliac disease. J Pediatr Gastroenterol Nutr. 2012; 54:136–160.
Article15. Adelman DC, Murray J, Wu TT, Mäki M, Green PH, Kelly CP. Measuring change in small intestinal histology in patients with celiac disease. Am J Gastroenterol. 2018; 113:339–347.
Article16. Elli L, Branchi F, Sidhu R, et al. Small bowel villous atrophy: celiac disease and beyond. Expert Rev Gastroenterol Hepatol. 2017; 11:125–138.
Article17. Singh P, Kurray L, Agnihotri A, et al. Titers of anti-tissue transglutaminase antibody correlate well with severity of villous abnormalities in celiac disease. J Clin Gastroenterol. 2015; 49:212–217.
Article18. Rahmati A, Shakeri R, Sohrabi M, et al. Correlation of tissue transglutaminase antibody with duodenal histologic marsh grading. Middle East J Dig Dis. 2014; 6:131–136.19. Hammer ST, Greenson JK. The clinical significance of duodenal lymphocytosis with normal villus architecture. Arch Pathol Lab Med. 2013; 137:1216–1219.
Article20. Trejdosiewicz LK. What is the role of human intestinal intraepithelial lymphocytes? Clin Exp Immunol. 1993; 94:395–397.
Article21. Veress B, Franzén L, Bodin L, Borch K. Duodenal intraepithelial lymphocyte-count revisited. Scand J Gastroenterol. 2004; 39:138–144.
Article22. Järvinen TT, Collin P, Rasmussen M, et al. Villous tip intraepithelial lymphocytes as markers of early-stage coeliac disease. Scand J Gastroenterol. 2004; 39:428–433.
Article23. Shalimar DM, Das P, Sreenivas V, Gupta SD, Panda SK, Makharia GK. Mechanism of villous atrophy in celiac disease: role of apoptosis and epithelial regeneration. Arch Pathol Lab Med. 2013; 137:1262–1269.
Article24. Ghosal UC, Das P. Diagnosis of celiac disease. In : Makharia GK, editor. Handbook of celiac disease. 1st ed. New Delhi: Kontentworx Publications;2015. p. 63–85.25. Švajdler M, Daum O, Rychlý B. Diagnosing celiac disease: role of the pathologists. Int J Celiac Dis. 2014; 2:70–75.
Article26. Rosekrans PC, Meijer CJ, Polanco I, Mearin ML, van der Wal AM, Lindeman J. Long-term morphological and immunohistochemical observations on biopsy specimens of small intestine from children with gluten-sensitive enteropathy. J Clin Pathol. 1981; 34:138–144.
Article27. Boruah D, Bhatia JK, Kamal KD, Malik A. Morphometric changes in jejunal mucosa in HIV positive patients presenting with enteropathy: an Indian study. Ann Pathol Lab Med. 2017; 4:A379–A387.
Article28. Hegenbart S, Uhl A, Vécsei A. Survey on computer aided decision support for diagnosis of celiac disease. Comput Biol Med. 2015; 65:348–358.
Article29. Hagendorn E, Whitney-Miller C, Huber A, Potts SJ. Development of a tissue image analysis algorithm for celiac drug development. In : Potts SJ, Eberhard DA, Wharton KA, editors. Methods in pharmacology and toxicology. New York: Springer;2015. p. 141–152.30. Vécsei A, Amann G, Hegenbart S, Liedlgruber M, Uhl A. Automated Marsh-like classification of celiac disease in children using local texture operators. Comput Biol Med. 2011; 41:313–325.31. Ciaccio EJ, Tennyson CA, Lewis SK, Krishnareddy S, Bhagat G, Green PH. Distinguishing patients with celiac disease by quantitative analysis of videocapsule endoscopy images. Comput Methods Programs Biomed. 2010; 100:39–48.
Article32. Gottlieb K, Dawson J, Hussain F, Murray JA. Development of drugs for celiac disease: review of endpoints for Phase 2 and 3 trials. Gastroenterol Rep (Oxf). 2015; 3:91–102.
Article33. Murray JA, Kelly CP, Green PH, et al. No difference between latiglutenase and placebo in reducing villous atrophy or improving symptoms in patients with symptomatic celiac disease. Gastroenterology. 2017; 152:787–798.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Celiac Disease: A Disorder Emerging from Antiquity, Its Evolving Classification and Risk, and Potential New Treatment Paradigms
- Comparison of Endoscopic and Histological Findings between Typical and Atypical Celiac Disease in Children
- A simple phenotypic classification for celiac disease
- Adult Celiac Disease and Its Malignant Complications
- A case of enteropathy-type intestinal T-cell lymphoma, confused with celiac disease