Ann Lab Med.
2018 May;38(3):226-234. 10.3343/alm.2018.38.3.226.
Comparison of Enterococcus faecium Bacteremic Isolates from Hematologic and Non-hematologic Patients: Differences in Antimicrobial Resistance and Molecular Characteristics
- Affiliations
-
- 1Division of Infectious Diseases, Department of Internal Medicine, Seoul St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea.
- 2Department of Laboratory Medicine, Seoul St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea. yjpk@catholic.ac.kr
- 3Department of Brain and Cognitive Science, Massachusetts Institute of Technology, Massachusetts, United States of America.
- 4Vaccine Bio Research Institute, Seoul St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea.
- 5Catholic Blood and Marrow Transplantation Center, Seoul St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea.
- KMID: 2403456
- DOI: http://doi.org/10.3343/alm.2018.38.3.226
Abstract
- BACKGROUND
Enterococcus faecium, especially vancomycin-resistant E. faecium (VREfm), is a major concern for patients with hematologic diseases. Exposure to antibiotics including fluoroquinolone, which is used as a routine prophylaxis for patients with hematologic (MH) diseases, has been reported to be a risk factor for infection with vancomycin-resistant eneterocci. We compared the characteristics of E. faecium isolates according to their vancomycin susceptibility and patient group (MH vs non-MH patients).
METHODS
A total of 120 E. faecium bacteremic isolates (84 from MH and 36 from non-MH patients) were collected consecutively, and their characteristics (susceptibility, multilocus sequence type [MLST], Tn1546 type, and the presence of virulence genes and plasmids) were determined.
RESULTS
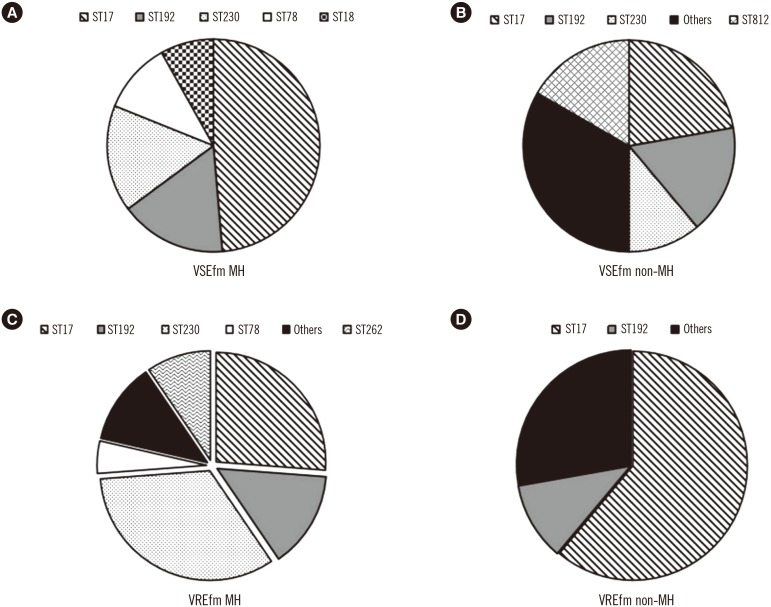
Among the vancomycin-susceptible E. faecium (VSEfm) isolates, resistance to ampicillin (97.6% vs 61.1%) and high-level gentamicin (71.4% vs 38.9%) was significantly higher in isolates from MH patients than in those from non-MH patients. Notably, hyl, esp, and pEF1071 were present only in isolates with ampicillin resistance. Among the VREfm isolates, ST230 (33.3%) and ST17 (26.2%) were predominant in MH patients, while ST17 (61.1%) was predominant in non-MH patients. Plasmid pLG1 was more prevalent in E. faecium isolates from MH patients than in those from non-MH patients, regardless of vancomycin resistance. Transposon analysis revealed five types across all VREfm isolates.
CONCLUSIONS
The antimicrobial resistance profiles and molecular characteristics of E. faecium isolates differed according to the underlying diseases of patients within the same hospital. We hypothesize that the prophylactic use of fluoroquinolone might have an effect on these differences.
Keyword
MeSH Terms
Figure
Reference
-
1. Wisplinghoff H, Bischoff T, Tallent SM, Seifert H, Wenzel RP, Edmond MB. Nosocomial bloodstream infections in US hospitals: analysis of 24,179 cases from a prospective nationwide surveillance study. Clin Infect Dis. 2004; 39:309–317. PMID: 15306996.2. Cho SY, Lee DG, Choi SM, Kwon JC, Kim SH, Choi JK, et al. Impact of vancomycin resistance on mortality in neutropenic patients with enterococcal bloodstream infection: a retrospective study. BMC Infect Dis. 2013; 13:504. PMID: 24164924.3. Kwon JC, Kim SH, Choi JK, Cho SY, Park YJ, Park SH, et al. Epidemiology and clinical features of bloodstream infections in hematology wards: one year experience at the Catholic Blood and Marrow Transplantation Center. Infect Chemother. 2013; 45:51–61. PMID: 24265950.4. Vergis EN, Hayden MK, Chow JW, Snydman DR, Zervos MJ, Linden PK, et al. Determinants of vancomycin resistance and mortality rates in enterococcal bacteremia. a prospective multicenter study. Ann Intern Med. 2001; 135:484–492. PMID: 11578151.5. Peel T, Cheng AC, Spelman T, Huysmans M, Spelman D. Differing risk factors for vancomycin-resistant and vancomycin-sensitive enterococcal bacteraemia. Clin Microbiol Infect. 2012; 18:388–394. PMID: 21848977.6. Bonten MJ, Willems R, Weinstein RA. Vancomycin-resistant enterococci: why are they here, and where do they come from? Lancet Infect Dis. 2001; 1:314–325. PMID: 11871804.7. Rosvoll TC, Pedersen T, Sletvold H, Johnsen PJ, Sollid JE, Simonsen GS, et al. PCR-based plasmid typing in Enterococcus faecium strains reveals widely distributed pRE25-, pRUM-, pIP501- and pHTbeta-related replicons associated with glycopeptide resistance and stabilizing toxin-antitoxin systems. FEMS Immunol Med Microbiol. 2010; 58:254–268. PMID: 20015231.8. Cattoir V, Leclercq R. Twenty-five years of shared life with vancomycin-resistant enterococci: is it time to divorce? J Antimicrob Chemother. 2013; 68:731–742. PMID: 23208830.9. De Angelis G, Cataldo MA, De Waure C, Venturiello S, La Torre G, Cauda R, et al. Infection control and prevention measures to reduce the spread of vancomycin-resistant enterococci in hospitalized patients: a systematic review and meta-analysis. J Antimicrob Chemother. 2014; 69:1185–1192. PMID: 24458513.10. Garner JS, Jarvis WR, Emori TG, Horan TC, Hughes JM. CDC definitions for nosocomial infections. Am J Infect Control. 1988; 16:128–140. PMID: 2841893.11. CLSI. Performance standards for antimicrobial susceptibility testing. 24th ed. CLSI supplement, M100-S24. Wayne, PA: Clinical and Laboratory Standards Institute;2014.12. Homan WL, Tribe D, Poznanski S, Li M, Hogg G, Spalburg E, et al. Multilocus sequence typing scheme for Enterococcus faecium. J Clin Microbiol. 2002; 40:1963–1971. PMID: 12037049.13. Rosvoll TC, Lindstad BL, Lunde TM, Hegstad K, Aasnaes B, Hammerum AM, et al. Increased high-level gentamicin resistance in invasive Enterococcus faecium is associated with aac(6′)Ie-aph(2′′)Ia-encoding transferable megaplasmids hosted by major hospital-adapted lineages. FEMS Immunol Med Microbiol. 2012; 66:166–176. PMID: 22672387.14. Vankerckhoven V, van Autgaerden T, Vael C, Lammens C, Chapelle S, Rossi R, et al. Development of a multiplex PCR for the detection of asa1, gelE, cylA, esp, and hyl genes in enterococci and survey for virulence determinants among European hospital isolates of Enterococcus faecium. J Clin Microbiol. 2004; 42:4473–4479. PMID: 15472296.15. Park SH, Park C, Choi SM, Lee DG, Kim SH, Kwon JC, et al. Molecular epidemiology of vancomycin-resistant Enterococcus faecium bloodstream infections among patients with neutropenia over a 6-year period in South Korea. Microb Drug Resist. 2011; 17:59–65. PMID: 21128837.16. Lee DG, Kim SH, Kim SY, Kim CJ, Park WB, Song YG, et al. Evidence-based guidelines for empirical therapy of neutropenic fever in Korea. Korean J Intern Med. 2011; 26:220–252. PMID: 21716917.17. Weissinger F, Auner HW, Bertz H, Buchheidt D, Cornely OA, Egerer G, et al. Antimicrobial therapy of febrile complications after high-dose chemotherapy and autologous hematopoietic stem cell transplantation--guidelines of the Infectious Diseases Working Party (AGIHO) of the German Society of Hematology and Oncology (DGHO). Ann Hematol. 2012; 91:1161–1174. PMID: 22638755.18. Balla E, Dicks LM, Du Toit M, van der Merwe MJ, Holzapfel WH. Characterization and cloning of the genes encoding enterocin 1071A and enterocin 1071B, two antimicrobial peptides produced by Enterococcus faecalis BFE1071. Appl Environ Microbiol. 2000; 66:1298–1304. PMID: 10742203.19. Balla E, Dicks LM. Molecular analysis of the gene cluster involved in the production and secretion of enterocins 1071A and 1071B and of the genes responsible for the replication and transfer of plasmid pEF1071. Int J Food Microbiol. 2005; 99:33–45. PMID: 15718027.20. Dupre I, Zanetti S, Schito AM, Fadda G, Sechi LA. Incidence of virulence determinants in clinical Enteorococcus faecium and Enterococcus faecalis isolates collected in Sardinia (Italy). J Med Microbiol. 2003; 52:491–498. PMID: 12748268.21. Billstrom H, Ludn B, Sullivan A, Nord CE. Virulence and antimicrobial resistance in clinical Enterococcus faecium. Int J Antimicrob Agents. 2008; 32:374–377. PMID: 18715765.22. Sánchez-Díaz AM, Cuartero C, Rodríguez JD, Lozano S, Alonso JM, Rodríguez-Domínguez M, et al. The rise of ampicillin-resistant Enterococcus faecium high-risk clones as a frequent intestinal colonizer in oncohaematological neutropenic patients on levofloxacin prophylaxis: a risk for bacteraemia? Clin Microbiol Infect. 2016; 22:59.e1–59.e8. PMID: 26321668.23. Laverde Gomez JA, van Schaik W, Freitas AR, Coque TM, Weaver KE, Francia MV, et al. A multiresistance megaplasmid pLG1 bearing a hylEfm genomic island in hospital Enterococcus faecium isolates. Int J Med Microbiol. 2011; 301:165–175. PMID: 20951641.24. Yang J, Jiang Y, Guo L, Ye L, Ma Y, Luo Y. Prevalence of diverse clones of vancomycin-resistant Enterococcus faecium ST78 in a Chinese hospital. Microb Drug Resist. 2016; 22:294–300. PMID: 26652286.25. Rice LB, Lakticová V, Carias LL, Rudin S, Hutton R, Marshall SH. Transferable capacity for gastrointestinal colonization in Enterococcus faecium in a mouse model. J Infect Dis. 2009; 199:342–349. PMID: 19049434.26. van Wamel WJ, Hendrickx AP, Bonten MJ, Top J, Posthuma G, Willems RJ. Growth condition-dependent Esp expression by Enterococcus faecium affects initial adherence and biofilm formation. Infect Immun. 2007; 75:924–931. PMID: 17118984.27. Sava IG, Heikens E, Huebner J. Pathogenesis and immunity in enterococcal infections. Clin Microbiol Infect. 2010; 16:533–540. PMID: 20569264.28. Hendrickx AP, van Schaik W, Willems RJ. The cell wall architecture of Enterococcus faecium: from resistance to pathogenesis. Future Microbiol. 2013; 8:993–1010. PMID: 23902146.29. Willems RJ, Top J, van Schaik W, Leavis H, Bonten M, Sirén J, et al. Restricted gene flow among hospital subpopulations of Enterococcus faecium. MBio. 2012; 3:e00151–e00112. PMID: 22807567.30. Galloway-Peña JR, Nallapareddy SR, Arias CA, Eliopoulos GM, Murray BE. Analysis of clonality and antibiotic resistance among early clinical isolates of Enterococcus faecium in the United States. J Infect Dis. 2009; 200:1566–1573. PMID: 19821720.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Correlation between Infective Factors and Antibiotic Resistance in Enterococci Clinical Isolates in West of Iran
- Bloodstream Infections in Patients with Hematologic Diseases: Causative Organisms and Factors Associated with Resistance
- Susceptibility of Fosfomycin against Vancomycin Resistant Enterococci
- The Antimicrobial Resistance of Nosocomial Enterococcus faecium Infection in Patients Received Probiotics Containing Enterococcus faecium
- Evaluation of identification in Enterococcus Species by Using MicroScan(R)Panels