Ann Lab Med.
2018 Jan;38(1):17-22. 10.3343/alm.2018.38.1.17.
Extensively Drug-Resistant Escherichia coli Sequence Type 1642 Carrying an IncX3 Plasmid Containing the blaKPC-2 Gene Associated with Transposon Tn4401a
- Affiliations
-
- 1Department of Laboratory Medicine, Kosin University College of Medicine, Busan, Korea.
- 2Department of Laboratory Medicine and Research Institute of Bacterial Resistance, Yonsei University College of Medicine, Seoul, Korea. kscpjsh@yuhs.ac
- 3Department of Dental Hygiene, College of Medical and Life Science, Shilla University, Busan, Korea.
- 4Department of Laboratory Medicine, National Health Insurance Service Ilsan Hospital, Ilsan, Korea.
- KMID: 2397597
- DOI: http://doi.org/10.3343/alm.2018.38.1.17
Abstract
- BACKGROUND
Extensively drug-resistant (XDR) Enterobacteriaceae carrying the bla(KPC) gene have emerged as a major global therapeutic concern. The purpose of this study was to analyze the complete sequences of plasmids from KPC-2 carbapenemase-producing XDR Escherichia coli sequence type (ST) 1642 isolates.
METHODS
We performed antimicrobial susceptibility testing, PCR, multilocus sequence typing (MLST), and whole-genome sequencing to characterize the plasmid-mediated KPC-2-producing E. coli clinical isolates.
RESULTS
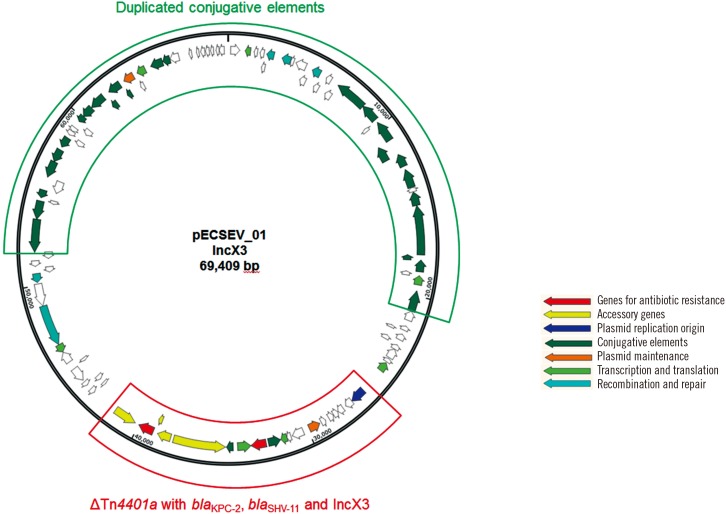
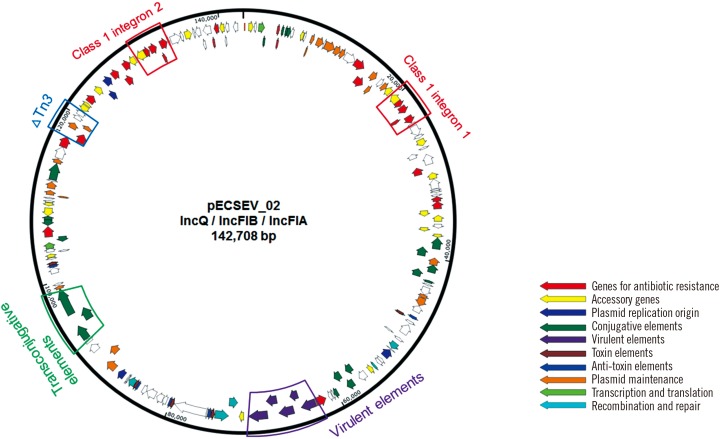
The isolates were resistant to most available antibiotics, including meropenem, ampicillin, ceftriaxone, gentamicin, and ciprofloxacin, but susceptible to tigecycline and colistin. The isolates were identified as the rare ST1642 by MLST. The isolates carried four plasmids: the first 69-kb conjugative IncX3 plasmid harbors bla(KPC-2) within a truncated Tn4401a transposon and bla(SHV-11) with duplicated conjugative elements. The second 142-kb plasmid with a multireplicon consisting of IncQ, IncFIA, and IncIB carries bla(TEM-1b) and two class 1 integrons. This plasmid also harbors a wide variety of additional antimicrobial resistance genes including aadA5, dfrA17, mph(A), sul1, tet(B), aac(3"²)-IId, strA, strB, and sul2.
CONCLUSIONS
The complete sequence analysis of plasmids from an XDR E. coli strain related to persistent infection showed the coexistence of a bla(KPC-2)-carrying IncX3-type plasmid and a class 1 integron-harboring multireplicon, suggesting its potential to cause outbreaks. Of additional clinical significance, the rare ST1642, identified in a cat, could constitute the source of human infection.
Keyword
MeSH Terms
-
Ampicillin
Animals
Anti-Bacterial Agents
Cats
Ceftriaxone
Ciprofloxacin
Colistin
Disease Outbreaks
Enterobacteriaceae
Escherichia coli*
Escherichia*
Gentamicins
Humans
Integrons
Multilocus Sequence Typing
Plasmids*
Polymerase Chain Reaction
Sequence Analysis
Ampicillin
Anti-Bacterial Agents
Ceftriaxone
Ciprofloxacin
Colistin
Gentamicins
Figure
Reference
-
1. Lee CR, Lee JH, Park KS, Kim YB, Jeong BC, Lee SH. Global dissemination of carbapenemase-producing Klebsiella pneumoniae: epidemiology, genetic Context, treatment options, and detection methods. Front Microbiol. 2016; 7:895. PMID: 27379038.2. Chen YT, Lin JC, Fung CP, Lu PL, Chuang YC, Wu TL, et al. KPC-2-encoding plasmids from Escherichia coli and Klebsiella pneumoniae in Taiwan. J Antimicrob Chemother. 2014; 69:628–631. PMID: 24123430.3. Adler A, Miller-Roll T, Assous MV, Geffen Y, Paikin S, Schwartz D, et al. A multicenter study of the clonal structure and resistance mechanism of KPC-producing Escherichia coli isolates in Israel. Clin Microbiol Infect. 2015; 21:230–235. PMID: 25658543.4. Piazza A, Caltagirone M, Bitar I, Nucleo E, Spalla M, Fogato E, et al. Emergence of Escherichia coli Sequence Type 131 (ST131) and ST3948 with KPC-2, KPC-3 and KPC-8 carbapenemases from a Long-Term Care and Rehabilitation Facility (LTCRF) in Northern Italy. Adv Exp Med Biol. 2016; 901:77–89. PMID: 26810233.5. Chavda KD, Chen L, Jacobs MR, Bonomo RA, Kreiswirth BN. Molecular diversity and plasmid analysis of KPC-producing Escherichia coli. Antimicrob Agents Chemother. 2016; 60:4073–4081. PMID: 27114279.6. Xu G, Jiang Y, An W, Wang H, Zhang X. Emergence of KPC-2-producing Escherichia coli isolates in an urban river in Harbin, China. World J Microbiol Biotechnol. 2015; 31:1443–1450. PMID: 26149956.7. Johnson TJ, Bielak EM, Fortini D, Hansen LH, Hasman H, Debroy C, et al. Expansion of the IncX plasmid family for improved identification and typing of novel plasmids in drug-resistant Enterobacteriaceae. Plasmid. 2012; 68:43–50. PMID: 22470007.8. He S, Chandler M, Varani AM, Hickman AB, Dekker JP, Dyda F. Mechanisms of evolution in high-consequence drug resistance plasmids. MBio. 2016; 7:e01987–e01916. PMID: 27923922.9. Naas T, Cuzon G, Villegas MV, Lartigue MF, Quinn JP, Nordmann P. Genetic structures at the origin of acquisition of the β-lactamase blaKPC gene. Antimicrob Agents Chemother. 2008; 52:1257–1263. PMID: 18227185.10. Naas T, Cuzon G, Truong HV, Nordmann P. Role of ISKpn7 and deletions in blaKPC gene expression. Antimicrob Agents Chemother. 2012; 56:4753–4759. PMID: 22733068.11. National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial susceptibility testing. Twenty-sixth informational supplement, M100-S26. Wayne, PA: National Committee for Clinical Laboratory Standards;2016.12. Kim MN, Yong D, An D, Chung HS, Woo JH, Lee K, et al. Nosocomial clustering of NDM-1-producing Klebsiella pneumoniae sequence type 340 strains in four patients at a South Korean tertiary care hospital. J Clin Microbiol. 2012; 50:1433–1436. PMID: 22259206.13. Chen L, Mediavilla JR, Endimiani A, Rosenthal ME, Zhao Y, Bonomo RA, et al. Multiplex real-time PCR assay for detection and classification of Klebsiella pneumoniae carbapenemase gene (blaKPC) variants. J Clin Microbiol. 2011; 49:579–585. PMID: 21123529.14. Jeong S, Kim JO, Jeong SH, Bae IK, Song W. Evaluation of peptide nucleic acid-mediated multiplex real-time PCR kits for rapid detection of carbapenemase genes in gram-negative clinical isolates. J Microbiol Methods. 2015; 113:4–9. PMID: 25819308.15. Jeong SH, Lee KM, Lee J, Bae IK, Kim JS, Kim HS, et al. Clonal and horizontal spread of the blaOXA-232 gene among Enterobacteriaceae in a Korean hospital. Diagn Microbiol Infect Dis. 2015; 82:70–72. PMID: 25702524.16. Harada K, Nakai Y, Kataoka Y. Mechanisms of resistance to cephalosporin and emergence of O25b-ST131 clone harboring CTX-M-27 β-lactamase in extraintestinal pathogenic Escherichia coli from dogs and cats in Japan. Microbiol Immunol. 2012; 56:480–485. PMID: 22486529.17. O'Hara JA, Hu F, Ahn C, Nelson J, Rivera JI, Pasculle AW, et al. Molecular epidemiology of KPC-producing Escherichia coli: occurrence of ST131-fimH30 subclone harboring pKpQIL-like IncFIIk plasmid. Antimicrob Agents Chemother. 2014; 58:4234–4237. PMID: 24820082.18. Kim JO, Song SA, Yoon EJ, Shin JH, Lee H, Jeong SH, et al. Outbreak of KPC-2-producing Enterobacteriaceae caused by clonal dissemination of Klebsiella pneumoniae ST307 carrying an IncX3-type plasmid harboring a truncated Tn4401a. Diagn Microbiol Infect Dis. 2017; 87:343–348. PMID: 28185686.19. Kassis-Chikhani N, Frangeul L, Drieux L, Sengelin C, Jarlier V, Brisse S, et al. Complete nucleotide sequence of the first KPC-2- and SHV-12-encoding IncX plasmid, pKpS90, from Klebsiella pneumoniae. Antimicrob Agents Chemother. 2013; 57:618–620. PMID: 23089759.20. Plante I, Centrón D, Roy PH. Direct sequencing and PCR mapping of integrons reveals multiple class 1 integrons in the multiresistant strain Enterobacter cloacae SCH88040794. FEMS Microbiol Lett. 2003; 221:59–62. PMID: 12694911.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Prevalence and molecular characteristics of carbapenemresistant Escherichia coli isolated from dogs in South Korea
- The expression of plasmid mediated afimbrial adhesin genes in an avian septicemic Escherichia coli strain
- Colistin resistance and plasmidmediated mcr genes in Escherichia coli and Salmonella isolated from pigs, pig carcass and pork in Thailand, Lao PDR and Cambodia border provinces
- Prevalence of TEM- and SHV-type Beta-lactamase gene in Escherichia coli and Klebsiella pneumoniae in Korea
- Prevalence of Escherichia coli Carrying pks Islands in Bacteremia Patients