Korean J Gastroenterol.
2022 Mar;79(3):118-125. 10.4166/kjg.2022.011.
Usefulness of Stool Multiplex Polymerase Chain Reaction Assays in Patients with Acute Diarrhea
- Affiliations
-
- 1Departments of Internal Medicine, Seoul Paik Hospital, Inje University College of Medicine, Seoul, Korea
- 2Departments of Laboratory Medicine, Seoul Paik Hospital, Inje University College of Medicine, Seoul, Korea
- KMID: 2527479
- DOI: http://doi.org/10.4166/kjg.2022.011
Abstract
- Background/Aims
There is a recent increase in the use of stool multiplex PCR assay-based diagnostic tests in patients with acute diarrhea. We used multiplex PCR assays to analyze the distribution of diarrhea-causing bacteria and viruses, as well as the clinical features of patients with acute diarrhea.
Methods
We retrospectively reviewed stool specimens of inpatients complaining of acute diarrhea from October 2018 to July 2020. The stool specimens had been tested for bacteria and viruses using multiplex PCR assays.
Results
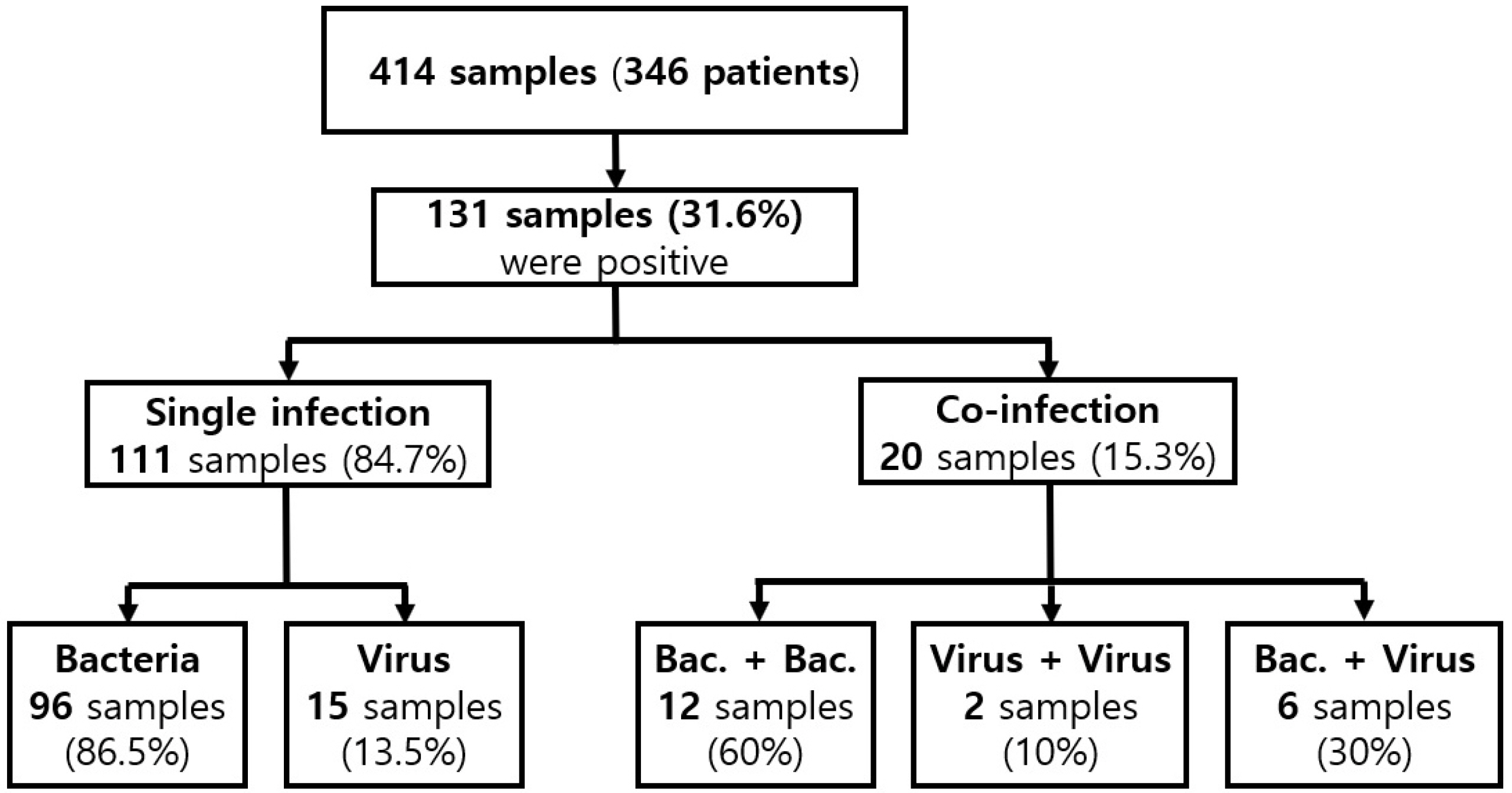
A total of 414 stool specimens from 346 patients were tested, and 152 pathogens were detected in 131 stool samples (131/414, 31.6%). Co-infection was detected in 20 patients (20/346, 5.8%). The common pathogens detected as causes of acute diarrhea, including co-infection, were Clostridium perfringens (34.9%), Clostridioides difficile (19.7%), and Campylobacter spp. (18.4%). The average age of patients with multiplex PCR-positive tests was lower than those with multiplex PCR-negative tests (p=0.001). In patients with suspected C. difficile infection (CDI), the RT-PCR for toxin gene assay was performed in 370 stool samples, 35 of which were positive (9.5%). Furthermore, 16 of the 35 samples were positive on the multiplex PCR assay (45.7%).
Conclusions
The multiplex PCR assay revealed that C. perfringens was the most common diarrhea-causing pathogen. In addition, in patients with suspected CDI, the multiplex PCR assay alone was insufficiently sensitive to detect pathogens and a conventional CDI test was additionally required.
Figure
Cited by 1 articles
-
Diagnostic value of serum procalcitonin and C-reactive protein in discriminating between bacterial and nonbacterial colitis: a retrospective study
Jae Yong Lee, So Yeon Lee, Yoo Jin Lee, Jin Wook Lee, Jeong Seok Kim, Ju Yup Lee, Byoung Kuk Jang, Woo Jin Chung, Kwang Bum Cho, Jae Seok Hwang
J Yeungnam Med Sci. 2023;40(4):388-393. doi: 10.12701/jyms.2023.00059.
Reference
-
1. WHO estimates of the global burden of foodborne diseases: foodborne disease burden epidemiology reference group 2007-2015. [Internet]. Geneva: World Health Organization;2015. cited 2021 Jul 1. Available from: https://apps.who.int/iris/handle/10665/199350.2. Shane AL, Mody RK, Crump JA, et al. 2017; 2017 Infectious Diseases Society of America clinical practice guidelines for the diagnosis and management of infectious diarrhea. Clin Infect Dis. 65:1963–1973. DOI: 10.1093/cid/cix959. PMID: 29194529. PMCID: PMC5848254.
Article3. DuPont HL. 2014; Acute infectious diarrhea in immunocompetent adults. N Engl J Med. 370:1532–1540. DOI: 10.1056/NEJMra1301069. PMID: 24738670.
Article4. Guerrant RL, Van Gilder T, Steiner TS, et al. 2001; Practice guidelines for the management of infectious diarrhea. Clin Infect Dis. 32:331–351. DOI: 10.1086/318514. PMID: 11170940.
Article5. Drancourt M. 2014; Multiplex testing of diarrhoea breaks down microbial barriers. Lancet Infect Dis. 14:663–664. DOI: 10.1016/S1473-3099(14)70844-8. PMID: 25022436.
Article6. Camilleri M. 2004; Chronic diarrhea: a review on pathophysiology and management for the clinical gastroenterologist. Clin Gastroenterol Hepatol. 2:198–206. DOI: 10.1016/s1542-3565(04)00003-5. PMID: 15017602.
Article7. Martín A, Pérez-Ayala A, Chaves F, Lora D, Orellana MÁ. 2018; Evaluation of the multiplex PCR Allplex-GI assay in the detection of bacterial pathogens in diarrheic stool samples. J Microbiol Methods. 144:33–36. DOI: 10.1016/j.mimet.2017.10.016. PMID: 29100990.
Article8. Owens RC Jr, Donskey CJ, Gaynes RP, Loo VG, Muto CA. 2008; Antimicrobial-associated risk factors for Clostridium difficile infection. Clin Infect Dis. 46(Suppl 1):S19–S31. DOI: 10.1086/521859. PMID: 18177218.9. Song PH, Min JH, Kim YS, et al. 2018; Rapid and accurate diagnosis of Clostridium difficile infection by real-time polymerase chain reaction. Intest Res. 16:109–115. DOI: 10.5217/ir.2018.16.1.109. PMID: 29422805. PMCID: PMC5797257.
Article10. Cho SH, Shin HH, Choi YH, Park MS, Lee BK. 2008; Enteric bacteria isolated from acute diarrheal patients in the Republic of Korea between the year 2004 and 2006. J Microbiol. 46:325–330. DOI: 10.1007/s12275-008-0015-4. PMID: 18604503.
Article11. Kim NO, Cha I, Kim JS, Chung GT, Kang YH, Hong S. 2013; The prevalence and characteristics of bacteria causing acute diarrhea in Korea, 2012. Ann Clin Microbiol. 16:174–181.
Article12. Bessède E, Delcamp A, Sifré E, Buissonnière A, Mégraud F. 2011; New methods for detection of campylobacters in stool samples in comparison to culture. J Clin Microbiol. 49:941–944. DOI: 10.1128/JCM.01489-10. PMID: 21209172. PMCID: PMC3067684.
Article13. Lee J, Kim J, Cho H, Oh K, Uh Y, Yoon KJ. 2015; Detection of bacterial and viral pathogens in stool specimens using multiplex PCR. Lab Med Qual Assur. 37:141–147.
Article14. Lee S, Park YJ, Lee HK, et al. 2013; Detection of 13 enteric bacteria and 5 viruses causing acute infectious diarrhea using multiplex PCR from direct stool specimens. Ann Clin Microbiol. 16:33–38.
Article15. Cho MC, Noh SA, Kim MN, Kim KM. 2010; Direct application of multiplex PCR on stool specimens for detection of enteropathogenic bacteria. Korean J Clin Microbiol. 13:162–168.
Article16. Kwack WG, Lim YJ, Kwon KH, Chung JW, Oh JY. 2020; Outcomes and clinical relevance of stool multiplex bacterial polymerase chain reaction in patients with acute diarrhea: single center experience. Korean J Intern Med. 35:300–309. DOI: 10.3904/kjim.2017.189. PMID: 30685964. PMCID: PMC7061005.
Article17. Park JO, Jeon JS, Kim JK. 2019; Epidemiologic trends of diarrhea-causing virus infection analyzed by multiplex reverse transcription PCR in Cheonan, Korea, 2010-2018. Microbiol Biotechnol Lett. 47:317–322.
Article18. Connor BA. 2018; Multiplex PCR testing for travelers' diarrhea-friend or foe? Int J Infect Dis. 66:96–98. DOI: 10.1016/j.ijid.2017.11.022. PMID: 29175449.
Article19. Freeman K, Tsertsvadze A, Taylor-Phillips S, et al. 2017; Agreement between gastrointestinal panel testing and standard microbiology methods for detecting pathogens in suspected infectious gastroenteritis: test evaluation and meta-analysis in the absence of a reference standard. PLoS One. 12:e0173196. DOI: 10.1371/journal.pone.0173196. PMID: 28253337. PMCID: PMC5333893.
Article20. Micic D, Hirsch A, Setia N, Rubin DT. 2018; Enteric infections complicating ulcerative colitis. Intest Res. 16:489–493. DOI: 10.5217/ir.2018.16.3.489. PMID: 30090049. PMCID: PMC6077301.
Article21. Grass JE, Gould LH, Mahon BE. 2013; Epidemiology of foodborne disease outbreaks caused by Clostridium perfringens, United States, 1998-2010. Foodborne Pathog Dis. 10:131–136. DOI: 10.1089/fpd.2012.1316. PMID: 23379281. PMCID: PMC4595929.
Article22. Scallan E, Hoekstra RM, Angulo FJ, et al. 2011; Foodborne illness acquired in the United States--major pathogens. Emerg Infect Dis. 17:7–15. DOI: 10.3201/eid1701.p11101. PMID: 21192848. PMCID: PMC3375761.
Article23. Doyle MP, Diez-Gonzalez F, Hill C. 2019. Food microbiology: fundamentals and frontiers. 5th ed. John Wiley & Sons;Hoboken (NJ):24. Juneja VK, Sofos JN. 2009. Pathogens and toxins in foods: challenges and interventions. ASM Press;Washington (DC):25. Viala C, Le Monnier A, Maataoui N, Rousseau C, Collignon A, Poilane I. 2012; Comparison of commercial molecular assays for toxigenic Clostridium difficile detection in stools: BD GeneOhm Cdiff, XPert C. difficile and illumigene C. difficile. J Microbiol Methods. 90:83–85. DOI: 10.1016/j.mimet.2012.04.017. PMID: 22565213.
Article26. Dekker JP, Frank KM. 2015; Salmonella, Shigella, and yersinia. Clin Lab Med. 35:225–246. DOI: 10.1016/j.cll.2015.02.002. PMID: 26004640. PMCID: PMC4443274.
Article27. Acheson D, Allos BM. 2001; Campylobacter jejuni infections: update on emerging issues and trends. Clin Infect Dis. 32:1201–1206. DOI: 10.1086/319760. PMID: 11283810.
Article28. Kim H, Lee SW, Chung SP, Kim SW, Yoo IS, Ha YR. 2002; Stool white-cell count as a predictor of long-term admission in healthy patients with acute diarrhea. J Korean Soc Emerg Med. 13:381–384.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Outcomes and clinical relevance of stool multiplex bacterial polymerase chain reaction in patients with acute diarrhea: single center experience
- Clinical Significance of Fecal Lactoferrin and Multiplex Polymerase Chain Reaction in Patients with Acute Diarrhea
- Evaluation of a New Multiplex Real-Time PCR Assay for Detecting Gastroenteritis-Causing Viruses in Stool Samples
- Rapid Diagnosis of Duchenne Muscular Dystrophy DMD by Multiplex Polymerase Chain Reaction PCR using Uncultured Amniocytes
- Detection of Bacterial and Viral Pathogens in Stool Specimens Using Multiplex PCR