Ann Lab Med.
2016 Jul;36(4):325-334. 10.3343/alm.2016.36.4.325.
Identification of Acinetobacter Species Using Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry
- Affiliations
-
- 1Department of Laboratory Medicine, Kosin University College of Medicine, Busan, Korea.
- 2Brain Korea 21 PLUS Project for Medical Science, Yonsei University, Seoul, Korea.
- 3Department of Laboratory Medicine and Research Institute of Bacterial Resistance, Yonsei University College of Medicine, Seoul, Korea. kscpjsh@yuhs.ac
- 4Department of Dental Hygiene, College of Medical and Life Science, Shilla University, Busan, Korea.
- KMID: 2373552
- DOI: http://doi.org/10.3343/alm.2016.36.4.325
Abstract
- BACKGROUND
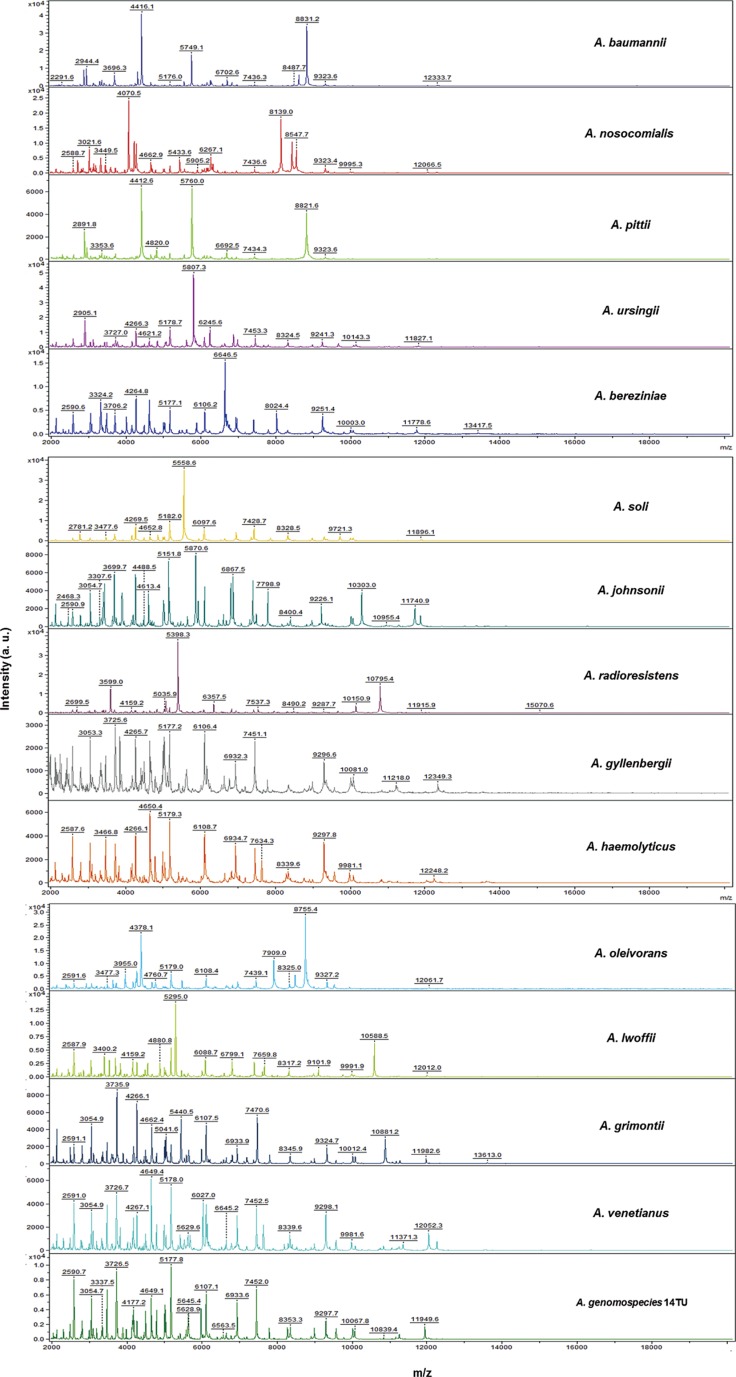
Acinetobacter baumannii has a greater clinical impact and exhibits higher antimicrobial resistance rates than the non-baumannii Acinetobacter species. Therefore, the correct identification of Acinetobacter species is clinically important. Matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry (MS) has recently become the method of choice for identifying bacterial species. The purpose of this study was to evaluate the ability of MALDI-TOF MS (Bruker Daltonics GmbH, Germany) in combination with an improved database to identify various Acinetobacter species.
METHODS
A total of 729 Acinetobacter clinical isolates were investigated, including 447 A. baumannii, 146 A. nosocomialis, 78 A. pittii, 18 A. ursingii, 9 A. bereziniae, 9 A. soli, 4 A. johnsonii, 4 A. radioresistens, 3 A. gyllenbergii, 3 A. haemolyticus, 2 A. lwoffii, 2 A. junii, 2 A. venetianus, and 2 A. genomospecies 14TU. After 212 isolates were tested with the default Bruker database, the profiles of 63 additional Acinetobacter strains were added to the default database, and 517 isolates from 32 hospitals were assayed for validation. All strains in this study were confirmed by rpoB sequencing.
RESULTS
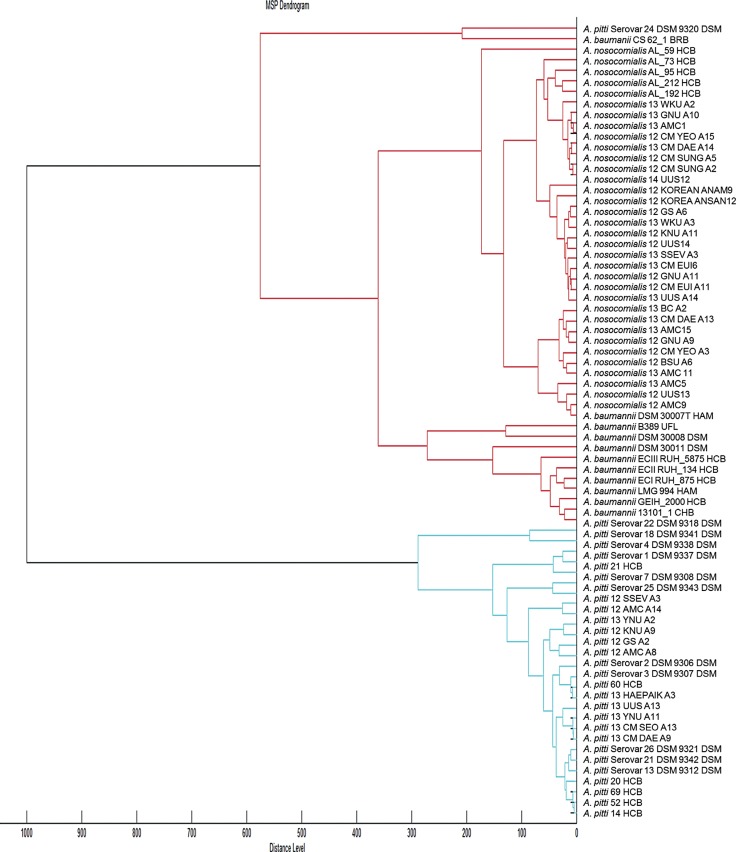
The addition of the 63 Acinetobacter strains' profiles to the default Bruker database increased the overall concordance rate between MALDI-TOF MS and rpoB sequencing from 69.8% (148/212) to 100.0% (517/517). Moreover, after library modification, all previously mismatched 64 Acinetobacter strains were correctly identified.
CONCLUSIONS
MALDI-TOF MS enables the prompt and accurate identification of clinically significant Acinetobacter species when used with the improved database.
Keyword
MeSH Terms
-
Acinetobacter Infections/*microbiology/pathology
Acinetobacter baumannii/*chemistry/classification/isolation & purification
Bacterial Proteins/chemistry/genetics/metabolism
Databases, Factual
Humans
Phylogeny
RNA, Ribosomal, 16S/chemistry/genetics/metabolism
*Spectrometry, Mass, Matrix-Assisted Laser Desorption-Ionization
Bacterial Proteins
RNA, Ribosomal, 16S
Figure
Cited by 2 articles
-
Colonization Prevalence and Risk Factor Analysis of Carbapenem-Resistant Acinetobacter baumannii in an Intensive Care Unit without Outbreaks
Young Ah Kim, Yoon Soo Park, Sang Sun Lee, Young Jun Son, Jeong Hwa Yeon, Young Hee Seo, Kyungwon Lee
Korean J Healthc Assoc Infect Control Prev. 2019;24(2):81-87. doi: 10.14192/kjicp.2019.24.2.81.새로운 MALDI-TOF질량분석기 ASTA MicroIDSys의
Acinetobacter 종 동정에 관한 성능 평가: Bruker Biotyper와의 비
Wooshin Kim, Ho Jong Lee, Seulbi Lee, Choon-Mee Kim, Jong Hee Shin, Soo Hyun Kim, Young Jin Ko, Seong-Ho Kang, Geon Park, Sook Jin Jang
Lab Med Online. 2022;12(3):183-190. doi: 10.47429/lmo.2022.12.3.183.
Reference
-
1. Espinal P, Roca I, Vila J. Clinical impact and molecular basis of antimicrobial resistance in non-baumannii Acinetobacter. Future Microbiol. 2011; 6:495–511. PMID: 21585259.
Article2. Kishii K, Kikuchi K, Matsuda N, Yoshida A, Okuzumi K, Uetera Y, et al. Evaluation of matrix-assisted laser desorption ionization-time of flight mass spectrometry for species identification of Acinetobacter strains isolated from blood cultures. Clin Microbiol Infect. 2014; 20:424–430. PMID: 24125498.
Article3. Lockhart SR, Abramson MA, Beekmann SE, Gallagher G, Riedel S, Diekema DJ, et al. Antimicrobial resistance among Gram-negative bacilli causing infections in intensive care unit patients in the United States between 1993 and 2004. J Clin Microbiol. 2007; 45:3352–3359. PMID: 17715376.
Article4. Kempf M, Rolain JM. Emergence of resistance to carbapenems in Acinetobacter baumannii in Europe: clinical impact and therapeutic options. Int J Antimicrob Agents. 2012; 39:105–114. PMID: 22113193.
Article5. Fournier PE, Richet H. The epidemiology and control of Acinetobacter baumannii in health care facilities. Clin Infect Dis. 2006; 42:692–699. PMID: 16447117.
Article6. Chen SF, Chang WN, Lu CH, Chuang YC, Tsai HH, Tsai NW, et al. Adult Acinetobacter meningitis and its comparison with non-Acinetobacter gram-negative bacterial meningitis. Acta Neurol Taiwan. 2005; 14:131–137. PMID: 16252615.7. Tsai HY, Cheng A, Liu CY, Huang YT, Lee YC, Liao CH, et al. Bacteremia caused by Acinetobacter junii at a medical center in Taiwan, 2000-2010. Eur J Clin Microbiol Infect Dis. 2012; 31:2737–2743. PMID: 22562410.
Article8. Castellanos Martínez E, Telenti Asensio M, Rodríguez Blanco VM, Rodríguez Suárez ML, Morena Torrico A, Cortina Llosa A. Infective endocarditis of an interventricular patch caused by Acinetobacter haemolyticus. Infection. 1995; 23:243–245. PMID: 8522385.9. Molina J, Cisneros JM, Fernández-Cuenca F, Rodríguez-Baño J, Ribera A, Beceiro A, et al. Clinical features of infections and colonization by Acinetobacter genospecies 3. J Clin Microbiol. 2010; 48:4623–4626. PMID: 20943868.10. Wisplinghoff H, Edmond MB, Pfaller MA, Jones RN, Wenzel RP, Seifert H. Nosocomial bloodstream infections caused by Acinetobacter species in United States hospitals: clinical features, molecular epidemiology, and antimicrobial susceptibility. Clin Infect Dis. 2000; 31:690–697. PMID: 11017817.
Article11. Chuang YC, Sheng WH, Li SY, Lin YC, Wang JT, Chen YC, et al. Influence of genospecies of Acinetobacter baumannii complex on clinical outcomes of patients with acinetobacter bacteremia. Clin Infect Dis. 2011; 52:352–360. PMID: 21193494.
Article12. Turton JF, Shah J, Ozongwu C, Pike R. Incidence of Acinetobacter species other than A. baumannii among clinical isolates of Acinetobacter: evidence for emerging species. J Clin Microbiol. 2010; 48:1445–1449. PMID: 20181894.13. Lee K, Yong D, Jeong SH, Chong Y. Multidrug-resistant Acinetobacter spp.: increasingly problematic nosocomial pathogens. Yonsei Med J. 2011; 52:879–891. PMID: 22028150.14. Wang J, Ruan Z, Feng Y, Fu Y, Jiang Y, Wang H, et al. Species distribution of clinical Acinetobacter isolates revealed by different identification techniques. PLoS One. 2014; 9:e104882. PMID: 25120020.
Article15. Chang HC, Wei YF, Dijkshoorn L, Vaneechoutte M, Tang CT, Chang TC. Species-level identification of isolates of the Acinetobacter calcoaceticus-Acinetobacter baumannii complex by sequence analysis of the 16S-23S rRNA gene spacer region. J Clin Microbiol. 2005; 43:1632–1639. PMID: 15814977.16. Gundi VA, Dijkshoorn L, Burignat S, Raoult D, La Scola B. Validation of partial rpoB gene sequence analysis for the identification of clinically important and emerging Acinetobacter species. Microbiology. 2009; 155:2333–2341. PMID: 19389786.
Article17. La Scola B, Gundi VA, Khamis A, Raoult D. Sequencing of the rpoB gene and flanking spacers for molecular identification of Acinetobacter species. J Clin Microbiol. 2006; 44:827–832. PMID: 16517861.18. Carbonnelle E, Mesquita C, Bille E, Day N, Dauphin B, Beretti JL, et al. MALDI-TOF mass spectrometry tools for bacterial identification in clinical microbiology laboratory. Clin Biochem. 2011; 44:104–109. PMID: 20620134.
Article19. Murray PR. What is new in clinical microbiology-microbial identification by MALDI-TOF mass spectrometry: a paper from the 2011 William Beaumont Hospital Symposium on molecular pathology. J Mol Diagn. 2012; 14:419–423. PMID: 22795961.20. Espinal P, Seifert H, Dijkshoorn L, Vila J, Roca I. Rapid and accurate identification of genomic species from the Acinetobacter baumannii (Ab) group by MALDI-TOF MS. Clin Microbiol Infect. 2012; 18:1097–1103. PMID: 22085042.
Article21. Jeong S, Kim JO, Jeong SH, Bae IK, Song W. Evaluation of peptide nucleic acid-mediated multiplex real-time PCR kits for rapid detection of carbapenemase genes in gram-negative clinical isolates. J Microbiol Methods. 2015; 113:4–9. PMID: 25819308.
Article22. Hsueh PR, Kuo LC, Chang TC, Lee TF, Teng SH, Chuang YC, et al. Evaluation of the Bruker Biotyper matrix-assisted laser desorption ionization-time of flight mass spectrometry system for identification of blood isolates of Acinetobacter species. J Clin Microbiol. 2014; 52:3095–3100. PMID: 24899038.
Article23. Šedo O, Nemec A, Křížová L, Kačalová M, Zdráhal Z. Improvement of MALDI-TOF MS profiling for the differentiation of species within the Acinetobacter calcoaceticus-Acinetobacter baumannii complex. Syst Appl Microbiol. 2013; 36:572–578. PMID: 24054697.
Article24. Donnarumma F, Sergi S, Indorato C, Mastromei G, Monnanni R, Nicoletti P, et al. Molecular characterization of acinetobacter isolates collected in intensive care units of six hospitals in Florence, Italy, during a 3-year surveillance program: a population structure analysis. J Clin Microbiol. 2010; 48:1297–1304. PMID: 20181903.25. Máder K, Terhes G, Hajdú E, Urbán E, Sóki J, Magyar T, et al. Outbreak of septicaemic cases caused by Acinetobacter ursingii in a neonatal intensive care unit. Int J Med Microbiol. 2010; 300:338–340. PMID: 19931486.
Article26. Idzenga D, Schouten MA, van Zanten AR. Outbreak of Acinetobacter genomic species 3 in a Dutch intensive care unit. J Hosp Infect. 2006; 63:485–487. PMID: 16815591.
Article27. van Dessel H, Kamp-Hopmans TE, Fluit AC, Brisse S, de Smet AM, Dijkshoorn L, et al. Outbreak of a susceptible strain of Acinetobacter species 13 (sensu Tjernberg and Ursing) in an adult neurosurgical intensive care unit. J Hosp Infect. 2002; 51:89–95. PMID: 12090795.
Article28. Park KH, Shin JH, Lee SY, Kim SH, Jang MO, Kang SJ, et al. The clinical characteristics, carbapenem resistance, and outcome of Acinetobacter bacteremia according to genospecies. PLoS One. 2013; 8:e65026. PMID: 23755171.
Article29. Lin YC, Sheng WH, Chen YC, Chang SC, Hsia KC, Li SY. Differences in carbapenem resistance genes among Acinetobacterbaumannii, Acinetobacter genospecies 3 and Acinetobacter genospecies 13TU in Taiwan. Int J Antimicrob Agents. 2010; 35:439–443. PMID: 20106635.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Reliability of Acinetobacter baumannii Identification with Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry
- Risk of inaccurate species identification by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry and of false carbapenem resistance by automated susceptibility analysis of Enterobacter spp.
- Evaluation of Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry-Based VITEK MS System for the Identification of Acinetobacter Species from Blood Cultures: Comparison with VITEK 2 and MicroScan Systems
- Comparison of a New Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry Platform, ASTA MicroIDSys, With Bruker Biotyper for Species Identification
- Comparison of the Bruker Biotyper and VITEK MS Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry Systems Using a Formic Acid Extraction Method to Identify Common and Uncommon Yeast Isolates