Endocrinol Metab.
2021 Jun;36(3):637-646. 10.3803/EnM.2021.962.
Association of Protein Z with Prediabetes and Type 2 Diabetes
- Affiliations
-
- 1Department of Internal Medicine, Keimyung University Dongsan Hospital, Keimyung University School of Medicine, Daegu, Korea
- 2Department of Physiology, Keimyung University School of Medicine, Daegu, Korea
- KMID: 2517654
- DOI: http://doi.org/10.3803/EnM.2021.962
Abstract
- Background
Type 2 diabetes mellitus (T2DM) is a progressive metabolic disease. Early detection of prediabetes is important to reduce the risk of T2DM. Some cytokines are known to be associated with T2DM. Therefore, we aimed to identify cytokines as novel biomarkers of glucose dysmetabolism.
Methods
The first stage of the study included 43 subjects (13 subjects with newly diagnosed T2DM, 13 with prediabetes, and 16 with normoglycemia) for cytokine microarray analysis. Blood samples of the subjects were assessed for 310 cytokines to identify potential indicators of prediabetes. The second stage included 142 subjects (36 subjects with T2DM, 35 with prediabetes, and 71 with normoglycemia) to validate the potential cytokines associated with prediabetes.
Results
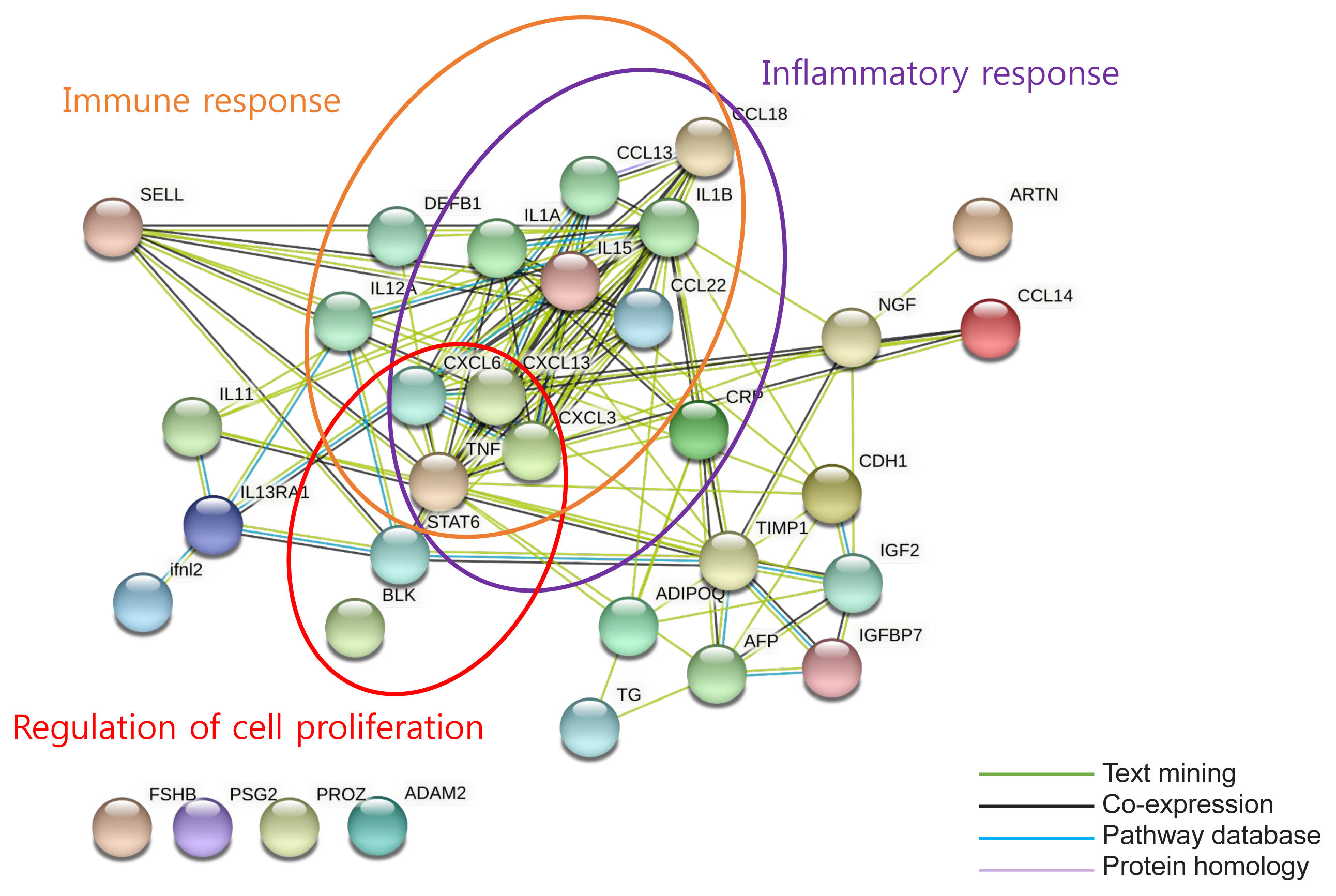
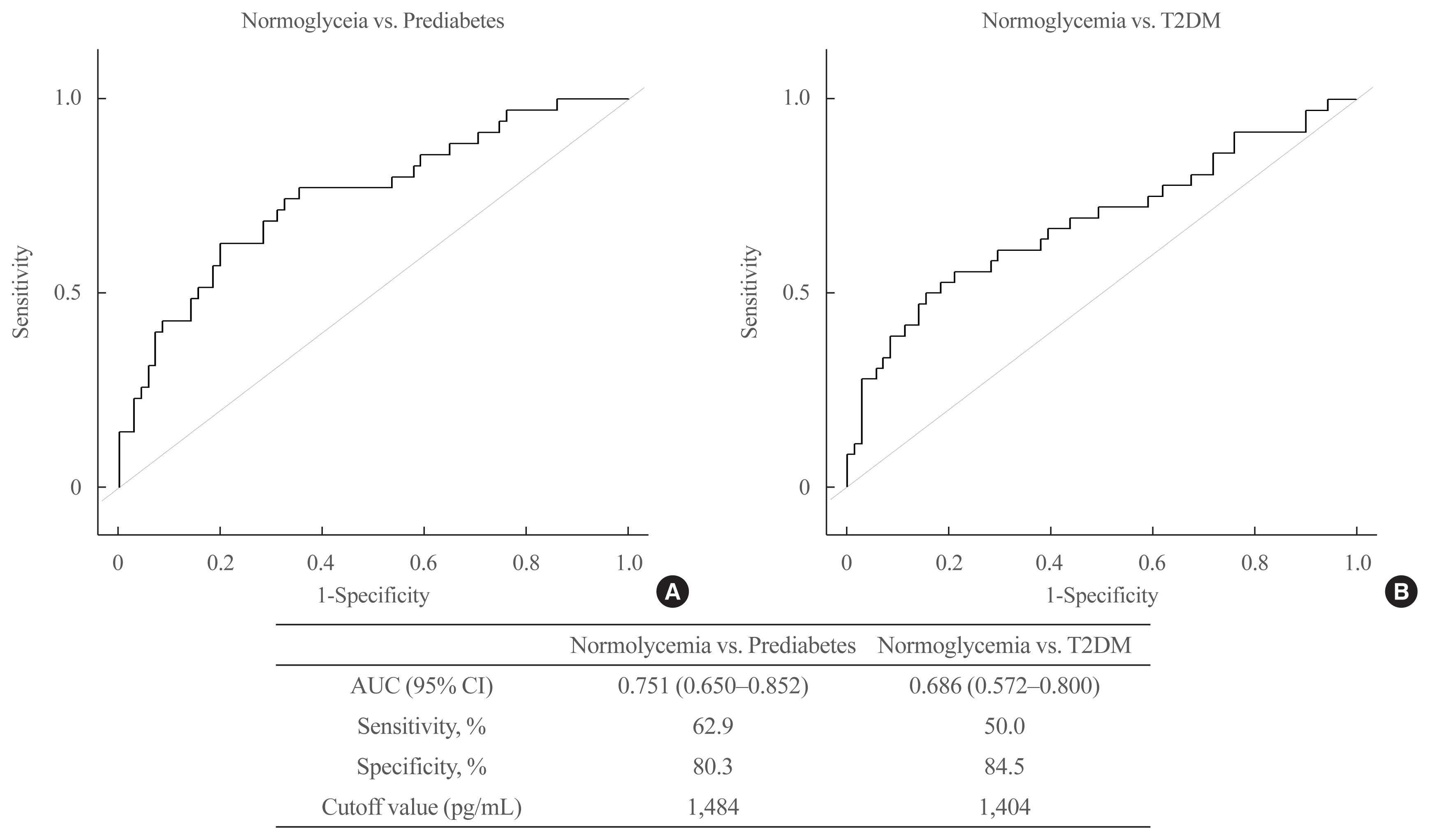
We identified 41 cytokines that differed by 1.5-fold or more in at least one out of the three comparisons (normoglycemia vs. prediabetes, normoglycemia vs. T2DM, and prediabetes vs. T2DM) among 310 cytokines. Finally, we selected protein Z (PROZ) and validated this finding to determine its association with prediabetes. Plasma PROZ levels were found to be decreased in patients with prediabetes (1,490.32±367.19 pg/mL) and T2DM (1,583.34±465.43 pg/mL) compared to those in subjects with normoglycemia (1,864.07±450.83 pg/mL) (P<0.001). There were significantly negative correlations between PROZ and fasting plasma glucose (P=0.001) and hemoglobin A1c (P=0.010).
Conclusion
PROZ levels were associated with prediabetes and T2DM. We suggest that PROZ may be a promising biomarker for the early detection of prediabetes. Further large-scale studies are needed to evaluate the relationship and mechanism between PROZ and prediabetes and T2DM.
Keyword
Figure
Cited by 3 articles
-
Identification of Protein Z as a Potential Novel Biomarker for the Diagnosis of Prediabetes
Seung-Hoi Koo
Endocrinol Metab. 2021;36(3):572-573. doi: 10.3803/EnM.2021.303.Association of Protein Z with Prediabetes and Type 2 Diabetes (
Endocrinol Metab 2021;36:637-46, Yun-Ui Bae et al.)
Tiffany Pascreau, Maia Tchikviladze, Emilie Jolly, Sara Zia-Chahabi, Bertrand Lapergue, Marc Vasse
Endocrinol Metab. 2021;36(5):1147-1148. doi: 10.3803/EnM.2021.1204.Association of Protein Z with Prediabetes and Type 2 Diabetes (
Endocrinol Metab 2021;36:637–46, Yun-Ui Bae et al.)
Ji Hong You, Yun-Ui Bae, Ho Chan Cho
Endocrinol Metab. 2021;36(5):1149-1150. doi: 10.3803/EnM.2021.502.
Reference
-
1. Tabak AG, Herder C, Rathmann W, Brunner EJ, Kivimaki M. Prediabetes: a high-risk state for diabetes development. Lancet. 2012; 379:2279–90.
Article2. American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care. 2010; 33(Suppl 1):S62–9.3. Hostalek U. Global epidemiology of prediabetes: present and future perspectives. Clin Diabetes Endocrinol. 2019; 5:5.4. Rask-Madsen C, King GL. Vascular complications of diabetes: mechanisms of injury and protective factors. Cell Metab. 2013; 17:20–33.
Article5. Milman S, Crandall JP. Mechanisms of vascular complications in prediabetes. Med Clin North Am. 2011; 95:309–25.
Article6. Bansal N. Prediabetes diagnosis and treatment: a review. World J Diabetes. 2015; 6:296–303.
Article7. Guo F, Moellering DR, Garvey WT. Use of HbA1c for diagnoses of diabetes and prediabetes: comparison with diagnoses based on fasting and 2-hr glucose values and effects of gender, race, and age. Metab Syndr Relat Disord. 2014; 12:258–68.
Article8. Lipska KJ, De Rekeneire N, Van Ness PH, Johnson KC, Kanaya A, Koster A, et al. Identifying dysglycemic states in older adults: implications of the emerging use of hemoglobin A1c. J Clin Endocrinol Metab. 2010; 95:5289–95.
Article9. Sehrawat T, Jindal A, Kohli P, Thour A, Kaur J, Sachdev A, et al. Utility and limitations of glycated hemoglobin (HbA1c) in patients with liver cirrhosis as compared with oral glucose tolerance test for diagnosis of diabetes. Diabetes Ther. 2018; 9:243–51.
Article10. Goldberg RB. Cytokine and cytokine-like inflammation markers, endothelial dysfunction, and imbalanced coagulation in development of diabetes and its complications. J Clin Endocrinol Metab. 2009; 94:3171–82.
Article11. Wang Z, Shen XH, Feng WM, Ye GF, Qiu W, Li B. Analysis of inflammatory mediators in prediabetes and newly diagnosed type 2 diabetes patients. J Diabetes Res. 2016; 2016:7965317.
Article12. Dorcely B, Katz K, Jagannathan R, Chiang SS, Oluwadare B, Goldberg IJ, et al. Novel biomarkers for prediabetes, diabetes, and associated complications. Diabetes Metab Syndr Obes. 2017; 10:345–61.
Article13. Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009; 4:44–57.
Article14. Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009; 37:1–13.
Article15. Doncheva NT, Morris JH, Gorodkin J, Jensen LJ. Cytoscape StringApp: network analysis and visualization of proteomics data. J Proteome Res. 2019; 18:623–32.
Article16. Broze GJ Jr, Miletich JP. Human protein Z. J Clin Invest. 1984; 73:933–8.
Article17. Han X, Fiehler R, Broze GJ Jr. Isolation of a protein Z-dependent plasma protease inhibitor. Proc Natl Acad Sci U S A. 1998; 95:9250–5.
Article18. Broze GJ Jr. Protein Z-dependent regulation of coagulation. Thromb Haemost. 2001; 86:8–13.
Article19. Fedi S, Sofi F, Brogi D, Tellini I, Cesari F, Sestini I, et al. Low protein Z plasma levels are independently associated with acute coronary syndromes. Thromb Haemost. 2003; 90:1173–8.
Article20. Heeb MJ, Paganini-Hill A, Griffin JH, Fisher M. Low protein Z levels and risk of ischemic stroke: differences by diabetic status and gender. Blood Cells Mol Dis. 2002; 29:139–44.
Article21. Santacroce R, Sarno M, Cappucci F, Sessa F, Colaizzo D, Brancaccio V, et al. Low protein Z levels and risk of occurrence of deep vein thrombosis. J Thromb Haemost. 2006; 4:2417–22.
Article22. Arora P, Garcia-Bailo B, Dastani Z, Brenner D, Villegas A, Malik S, et al. Genetic polymorphisms of innate immunity-related inflammatory pathways and their association with factors related to type 2 diabetes. BMC Med Genet. 2011; 12:95.
Article23. Donath MY, Shoelson SE. Type 2 diabetes as an inflammatory disease. Nat Rev Immunol. 2011; 11:98–107.
Article24. Maschirow L, Khalaf K, Al-Aubaidy HA, Jelinek HF. Inflammation, coagulation, endothelial dysfunction and oxidative stress in prediabetes: biomarkers as a possible tool for early disease detection for rural screening. Clin Biochem. 2015; 48:581–5.25. Berbudi A, Rahmadika N, Tjahjadi AI, Ruslami R. Type 2 diabetes and its impact on the immune system. Curr Diabetes Rev. 2020; 16:442–9.
Article26. Chen C, Cohrs CM, Stertmann J, Bozsak R, Speier S. Human beta cell mass and function in diabetes: recent advances in knowledge and technologies to understand disease pathogenesis. Mol Metab. 2017; 6:943–57.
Article27. Pradhan AD, Manson JE, Rifai N, Buring JE, Ridker PM. C-reactive protein, interleukin 6, and risk of developing type 2 diabetes mellitus. JAMA. 2001; 286:327–34.
Article28. Levi-Montalcini R, Skaper SD, Dal Toso R, Petrelli L, Leon A. Nerve growth factor: from neurotrophin to neurokine. Trends Neurosci. 1996; 19:514–20.
Article29. Calza L, Giardino L, Giuliani A, Aloe L, Levi-Montalcini R. Nerve growth factor control of neuronal expression of angiogenetic and vasoactive factors. Proc Natl Acad Sci U S A. 2001; 98:4160–5.
Article30. Cantarella G, Lempereur L, Presta M, Ribatti D, Lombardo G, Lazarovici P, et al. Nerve growth factor-endothelial cell interaction leads to angiogenesis in vitro and in vivo. FASEB J. 2002; 16:1307–9.
Article31. Park KS, Kim SS, Kim JC, Kim HC, Im YS, Ahn CW, et al. Serum and tear levels of nerve growth factor in diabetic retinopathy patients. Am J Ophthalmol. 2008; 145:432–7.
Article32. Vora SR, Zheng H, Stadler ZK, Fuchs CS, Zhu AX. Serum alpha-fetoprotein response as a surrogate for clinical outcome in patients receiving systemic therapy for advanced hepatocellular carcinoma. Oncologist. 2009; 14:717–25.33. Henriques CU, Damm P, Tabor A, Pedersen JF, Molsted-Pedersen L. Decreased alpha-fetoprotein in amniotic fluid and maternal serum in diabetic pregnancy. Obstet Gynecol. 1993; 82:960–4.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Sex differences in factors associated with prediabetes in Korean adults
- Family Involvement to Stop the Conversion of Prediabetes to Diabetes
- The effect of long working hours on developing type 2 diabetes in adults with prediabetes: The Kangbuk Samsung Cohort Study
- Prevention of Type 2 Diabetes
- Long-term association of pericardial adipose tissue with incident diabetes and prediabetes: the Coronary Artery Risk Development in Young Adults Study