J Vet Sci.
2017 Mar;18(1):51-58. 10.4142/jvs.2017.18.1.51.
Molecular-level evaluation of selected periodontal pathogens from subgingival regions in canines and humans with periodontal disease
- Affiliations
-
- 1Department and Clinic of Animal Surgery, Faculty of Veterinary Medicine, University of Life Sciences in Lublin, 20-612 Lublin, Poland. magdareba@wp.pl
- 2Dental Center ‘Lardent’, 20-135 Lublin, Poland.
- 3Children's Orthopaedic Clinic and Rehabilitation Department, Medical University of Lublin, 20-093 Lublin, Poland.
- KMID: 2412587
- DOI: http://doi.org/10.4142/jvs.2017.18.1.51
Abstract
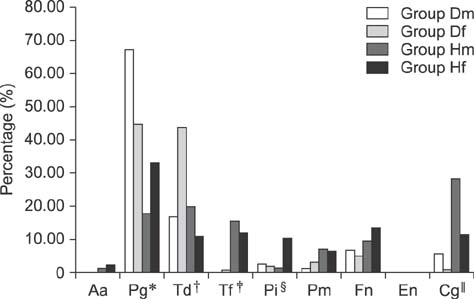
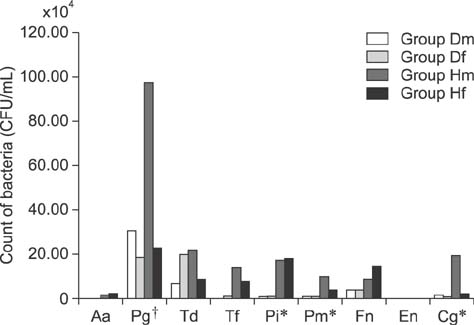
- Dogs commonly serve as a model for various human conditions, including periodontal diseases. The aim of this study was to identify the anaerobic bacteria that colonize the subgingival areas in dogs and humans by using rapid real-time polymerase chain reaction (RT-PCR)-based tests and to compare the results obtained in each species. Bacterial microflora evaluations, both quantitative and qualitative, were performed by applying ready-made tests on twelve dogs and twelve humans. Five samples were collected from each subject's deepest gingival pockets and joined to form a collective sample. The results of the study revealed interspecies similarities in the prevalences of Porphyromonas (P.) gingivalis, Treponema denticola, Tannerella forsythia, and Fusobacterium nucleatum. Red complex bacteria comprised the largest portion of the studied bacterial complexes in all study groups, with P. gingivalis being the most commonly isolated bacterium. The results show similarities in the prevalence of bacterial microflora in dogs and humans. Microbiological analysis of gingival pockets by using rapid real-time PCR-based tests in clinical practice, both veterinary and human, can facilitate the choice of appropriate pharmacological treatment and can provide a basis for subsequent verification of the treatment's effectiveness.
MeSH Terms
Figure
Reference
-
1. Abiko Y, Sato T, Mayanagi G, Takahashi N. Profiling of subgingival plaque biofilm microflora from periodontally healthy subjects and from subjects with periodontitis using quantitative real-time PCR. J Periodontal Res. 2010; 45:389–395.
Article2. Abusleme L, Dupuy AK, Dutzan N, Silva N, Burleson JA, Strausbaugh LD, Gamonal J, Diaz PI. The subgingival microbiome in health and periodontitis and its relationship with community biomass and inflammation. ISME J. 2013; 7:1016–1025.
Article3. Albandar JM. Aggressive and acute periodontal diseases. Periodontol 2000. 2014; 65:7–12.
Article4. Al-hebshi NN, Al-Alimi A, Taiyeb-Ali T, Jaafar N. Quantitative analysis of classical and new putative periodontal pathogens in subgingival biofilm: a case-control study. J Periodontal Res. 2015; 50:320–329.
Article5. Allaker RP, Young KA, Langlois T, de Rosayro R, Hardie JM. Dental plaque flora of the dog with reference to fastidious and anaerobic bacteria associated with bites. J Vet Dent. 1997; 14:127–130.
Article6. Boutaga K, van Winkelhoff AJ, Vandenbroucke-Grauls CMJE, Savelkoul PHM. Comparison of real-time PCR and culture for detection of Porphyromonas gingivalis in subgingival plaque samples. J Clin Microbiol. 2003; 41:4950–4954.
Article7. Boutaga K, van Winkelhoff AJ, Vandenbroucke-Grauls CMJE, Savelkoul PHM. Periodontal pathogens: a quantitative comparison of anaerobic culture and real-time PCR. FEMS Immunol Med Microbiol. 2005; 45:191–199.
Article8. Boutaga K, van Winkelhoff AJ, Vandenbroucke-Grauls CMJE, Savelkoul PHM. The additional value of real-time PCR in the quantitative detection of periodontal pathogens. J Clin Periodontol. 2006; 33:427–433.
Article9. Charalampakis G, Abrahamsson I, Carcuac O, Dahlén G, Berglundh T. Microbiota in experimental periodontitis and peri-implantitis in dogs. Clin Oral Implants Res. 2014; 25:1094–1098.
Article10. Colombo AP, Teles RP, Torres MC, Souto R, Rosalém W Jr, Mendes MCS, Uzeda M. Subgingival microbiota of Brazilian subjects with untreated chronic periodontitis. J Periodontol. 2002; 73:360–369.
Article11. da Silva-Boghossian CM, do Souto RM, Luiz RR, Colombo APV. Association of red complex, A. actinomycetemcomitans and non-oral bacteria with periodontal diseases. Arch Oral Biol. 2011; 56:899–906.
Article12. Dahlén G, Charalampakis G, Abrahamsson I, Bengtsson L, Falsen E. Predominant bacterial species in subgingival plaque in dogs. J Periodontal Res. 2012; 47:354–364.
Article13. Davis IJ, Wallis C, Deusch O, Colyer A, Milella L, Loman N, Harris S. A cross-sectional survey of bacterial species in plaque from client owned dogs with healthy gingiva, gingivitis or mild periodontitis. PLoS One. 2013; 8:e83158.
Article14. Dewhirst FE, Klein EA, Thompson EC, Blanton JM, Chen T, Milella L, Buckley CM, Davis IJ, Bennett ML, Marshall-Jones ZV. The canine oral microbiome. PLoS One. 2012; 7:e36067.
Article15. Di Bello A, Buonavoglia A, Franchini D, Valastro C, Ventrella G, Greco MF, Corrente M. Periodontal disease associated with red complex bacteria in dogs. J Small Anim Pract. 2014; 55:160–163.
Article16. Griffen AL, Beall CJ, Campbell JH, Firestone ND, Kumar PS, Yang ZK, Podar M, Leys EJ. Distinct and complex bacterial profiles in human periodontitis and health revealed by 16S pyrosequencing. ISME J. 2012; 6:1176–1185.
Article17. Haffajee AD, Teles RP, Socransky SS. Association of Eubacterium nodatum and Treponema denticola with human periodontitis lesions. Oral Microbiol Immunol. 2006; 21:269–282.
Article18. Hardham J, Dreier K, Wong J, Sfintescu C, Evans RT. Pigmented-anaerobic bacteria associated with canine periodontitis. Vet Microbiol. 2005; 106:119–128.
Article19. Hirai N, Shirai M, Kato Y, Murakami M, Nomura R, Yamasaki Y, Takahashi S, Kondo C, Matsumoto-Nakano M, Nakano K, Asai F. Correlation of age with distribution of periodontitis-related bacteria in Japanese dogs. J Vet Med Sci. 2013; 75:999–1001.
Article20. Kato Y, Shirai M, Murakami M, Mizusawa T, Hagimoto A, Wada K, Nomura R, Nakano K, Ooshima T, Asai F. Molecular detection of human periodontal pathogens in oral swab specimens from dogs in Japan. J Vet Dent. 2011; 28:84–89.
Article21. Khazandi M, Bird PS, Owens J, Wilson G, Meyer JN, Trott DJ. In vitro efficacy of cefovecin against anaerobic bacteria isolated from subgingival plaque of dogs and cats with periodontal disease. Anaerobe. 2014; 28:104–108.
Article22. Mane AK, Karmarkar AP, Bharadwaj RS. Anaerobic bacteria in subjects with chronic periodontitis and in periodontal health. J Oral Health Community Dent. 2009; 3:49–51.
Article23. Moore WE, Moore LV. The bacteria of periodontal diseases. Periodontol 2000. 1994; 5:66–77.
Article24. Mueller HP. Periodontology. Lublin: Thieme;2003. p. 15–29.25. Nakayama K. Porphyromonas gingivalis and related bacteria: from colonial pigmentation to the type IX secretion system and gliding motility. J Periodont Res. 2015; 50:1–8.
Article26. Nishiyama SAB, Senhorinho GNA, Gioso MA, Avila-Campos MJ. Detection of putative periodontal pathogens in subgingival specimens of dogs. Braz J Microbiol. 2007; 38:23–28.
Article27. Oz HS, Puleo DA. Animal models for periodontal disease. J Biomed Biotechnol. 2011; 2011:754857.
Article28. Papapanou PN, Neiderud AM, Papadimitriou A, Sandros J, Dahlén G. “Checkerboard” assessments of periodontal microbiota and serum antibody responses: a case-control study. J Periodontol. 2000; 71:885–897.
Article29. Paster BJ, Boches SK, Galvin JL, Ericson RE, Lau CN, Levanos VA, Sahasrabudhe A, Dewhirst FE. Bacterial diversity in human subgingival plaque. J Bacteriol. 2001; 183:3770–3783.
Article30. Raja M, Ummer F, Dhivakar CP. Aggregatibacter actinomycetemcomitans - a tooth killer? J Clin Diagn Res. 2014; 8:ZE13–ZE16.31. Riggio MP, Lennon A, Taylor DJ, Bennett D. Molecular identification of bacteria associated with canine periodontal disease. Vet Microbiol. 2011; 150:394–400.
Article32. Salari MH, Kadkhoda Z. Rate of cultivable subgingival periodontopathogenic bacteria in chronic periodontitis. J Oral Sci. 2004; 46:157–161.
Article33. Schacher B, Baron F, Rossberg M, Wohlfeil M, Arndt R, Eickholz P. Aggregatibacter actinomycetemcomitans as indicator for aggressive periodontitis by two analysing strategies. J Clin Periodontol. 2007; 34:566–573.
Article34. Senhorinho GN, Nakano V, Liu C, Song Y, Finegold SM, Avila-Campos MJ. Occurrence and antimicrobial susceptibility of Porphyromonas spp. and Fusobacterium spp. in dogs with and without periodontitis. Anaerobe. 2012; 18:381–385.
Article35. Socransky SS, Haffajee AD, Cugini MA, Smith C, Kent RL Jr. Microbial complexes in subgingival plaque. J Clin Periodontol. 1998; 25:134–144.
Article36. Socransky SS, Haffajee AD. Dental biofilms: difficult therapeutic targets. Periodontol 2000. 2002; 28:12–55.
Article37. Struillou X, Boutigny H, Soueidan A, Layrolle P. Experimental animal models in periodontology: a review. Open Dent J. 2010; 4:37–47.
Article38. Sturgeon A, Stull JW, Costa MC, Weese JS. Metagenomic analysis of the canine oral cavity as revealed by high-throughput pyrosequencing of the 16S rRNA gene. Vet Microbiol. 2013; 162:891–898.
Article39. Suzuki N, Yoshida A, Nakano Y. Quantitative analysis of multi-species oral biofilms by TaqMan Real-Time PCR. Clin Med Res. 2005; 3:176–185.
Article40. Yamasaki Y, Nomura R, Nakano K, Naka S, Matsumoto-Nakano M, Asai F, Ooshima T. Distribution of periodontopathic bacterial species in dogs and their owners. Arch Oral Biol. 2012; 57:1183–1188.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Relation between the interval of supportive periodontal therapy and the prevalence of the subgingival microflora
- Immediate effect of Nd:YAG laser monotherapy on subgingival periodontal pathogens: a pilot clinical study
- Efficacy of salivary versus subgingival bacterial sampling for the detection and quantification of periodontal pathogens
- Waiting for innovations in periodontal disease diagnosis
- Reply on "Relationship between maternal periodontal disease and Apgar score of newborns"