Int J Stem Cells.
2020 Nov;13(3):424-431. 10.15283/ijsc20048.
KLF4 Regulates Goblet Cell Differentiation in BMI1+ Reserve Intestinal Stem Cell Lineage during Homeostasis
- Affiliations
-
- 1Department of Medicine, Renaissance School of Medicine, Stony Brook University, Stony Brook, NY, USA
- 2Department of Gastroenterology and Metabolism, Nagoya City University Graduate School of Medical Sciences, Nagoya, Japan
- 3Department of Physiology and Biophysics, Renaissance School of Medicine, Stony Brook University, Stony Brook, NY, USA
- KMID: 2508916
- DOI: http://doi.org/10.15283/ijsc20048
Abstract
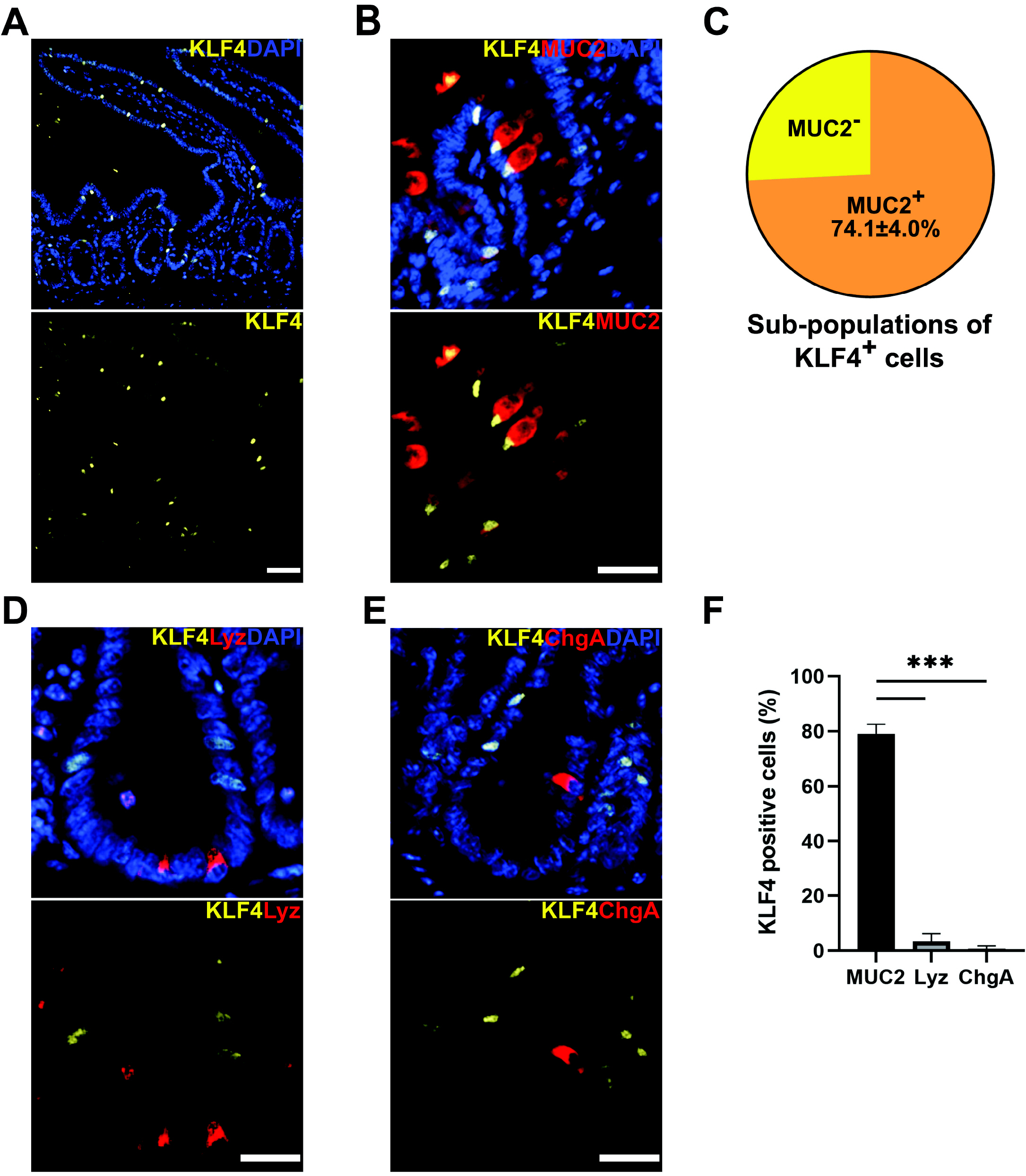
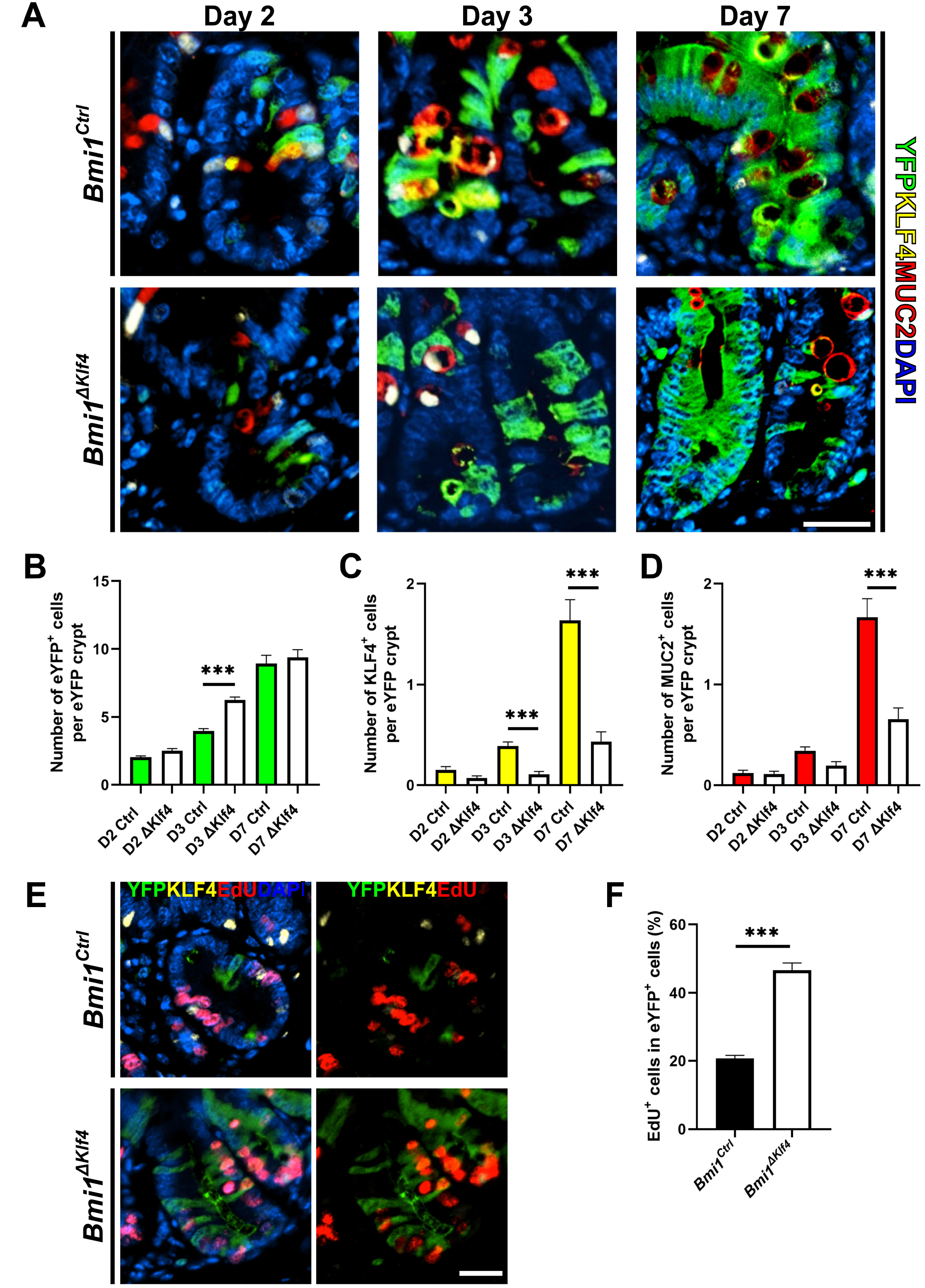
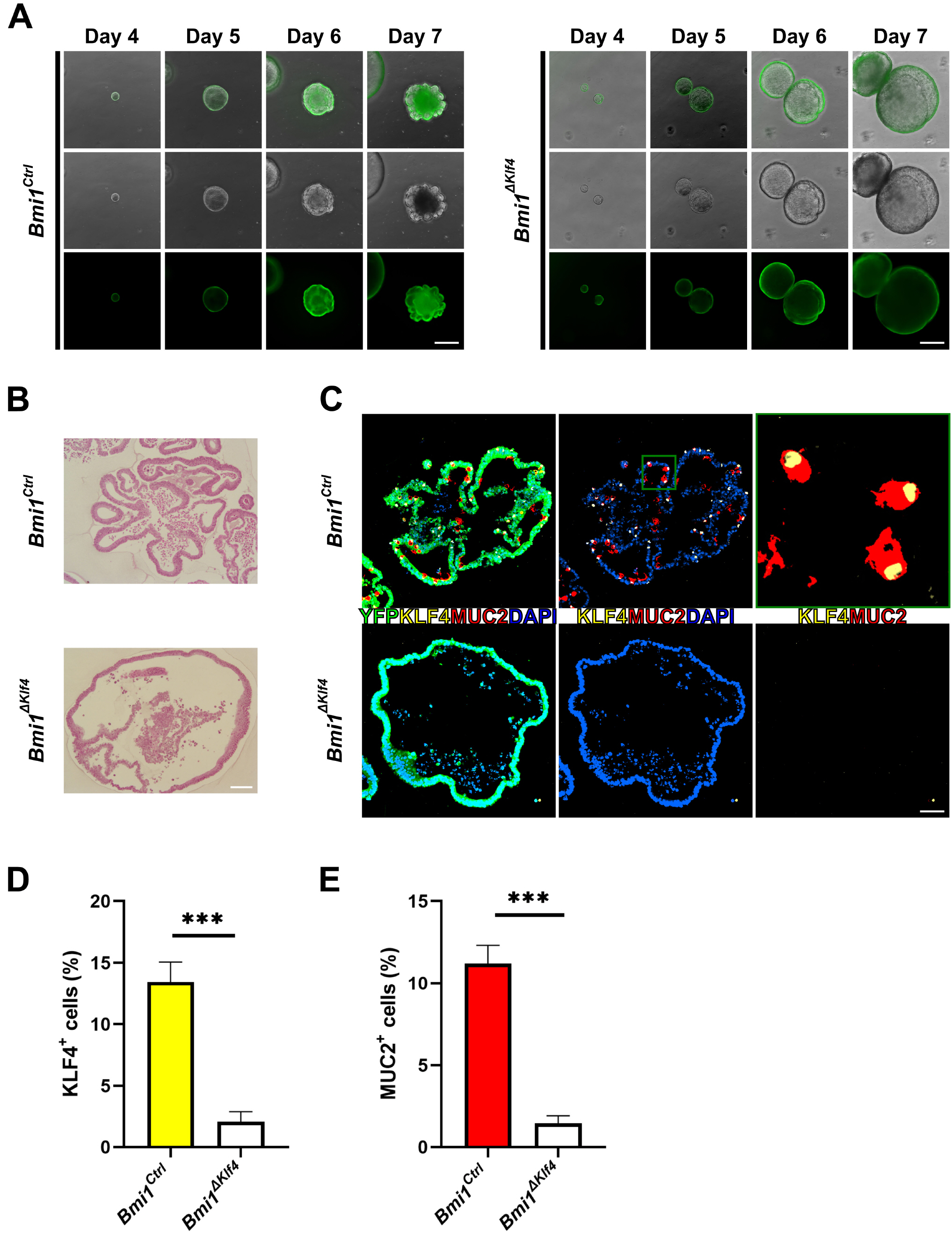
- Krüppel-like factor 4 (KLF4) is a zinc-finger transcription factor, expressed in villus cells of the intestinal epithelium, that promotes cellular differentiation and tissue homeostasis. Previous studies suggest that BMI1+ cells represent secretory progenitors with reserve intestinal stem cell (rISC) activity. However, it has not been elucidated how KLF4 contributes to crypt regeneration originated from BMI1+ rISC lineage during homeostasis. In this study, Bmi1-CreER ;Rosa26eYFP (Bmi1Ctrl ) and Bmi1-CreER ;Rosa26eYFP ;Klf4fl/fl (Bmi1△Klf4 ) mice were injected with tamoxifen to label BMI1+ cells and their lineage and to delete Klf4. During homeostasis, MUC2+ goblet cells appeared in the BMI1+ cell lineage 2, 3 and 7 days after tamoxifen administration. After Klf4 deletion in BMI1+ cells, the number of KLF4+ and MUC2+ cells in eYFP+ cells decreased in Bmi1△Klf4 mice compared with Bmi1Ctrl mice. Thus, KLF4 was positively correlated with goblet cell differentiation in BMI1+ cell derived lineage. In ex-vivo analysis, organoids derived from single eYFP+ cells of Bmi1Ctrl mice contained MUC2-expressing cells that co-expressed KLF4. On the other hand, organoids derived from Klf4-deleted eYFP+ cells from Bmi1△Klf4 mice showed reduced number of MUC2-expressing cells. In conclusion, these results suggest that KLF4 regulates goblet cell differentiation in BMI1+ ISC-derived lineage during homeostasis.
Figure
Reference
-
References
1. Barker N, van Es JH, Kuipers J, Kujala P, van den Born M, Cozijnsen M, Haegebarth A, Korving J, Begthel H, Peters PJ, Clevers H. 2007; Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature. 449:1003–1007. DOI: 10.1038/nature06196. PMID: 17934449.
Article2. Clevers H. 2013; The intestinal crypt, a prototype stem cell compartment. Cell. 154:274–284. DOI: 10.1016/j.cell.2013.07.004. PMID: 23870119.
Article3. Kim CK, Yang VW, Bialkowska AB. 2017; The role of intestinal stem cells in epithelial regeneration following radiation-induced gut injury. Curr Stem Cell Rep. 3:320–332. DOI: 10.1007/s40778-017-0103-7. PMID: 29497599. PMCID: PMC5818549.
Article4. Sangiorgi E, Capecchi MR. 2008; Bmi1 is expressed in vivo in intestinal stem cells. Nat Genet. 40:915–920. DOI: 10.1038/ng.165. PMID: 18536716. PMCID: PMC2906135.
Article5. Tian H, Biehs B, Warming S, Leong KG, Rangell L, Klein OD, de Sauvage FJ. 2011; A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature. 478:255–259. DOI: 10.1038/nature10408. PMID: 21927002. PMCID: PMC4251967.
Article6. Yan KS, Chia LA, Li X, Ootani A, Su J, Lee JY, Su N, Luo Y, Heilshorn SC, Amieva MR, Sangiorgi E, Capecchi MR, Kuo CJ. 2012; The intestinal stem cell markers Bmi1 and Lgr5 identify two functionally distinct populations. Proc Natl Acad Sci U S A. 109:466–471. DOI: 10.1073/pnas.1118857109. PMID: 22190486. PMCID: PMC3258636.
Article7. Noah TK, Donahue B, Shroyer NF. 2011; Intestinal development and differentiation. Exp Cell Res. 317:2702–2710. DOI: 10.1016/j.yexcr.2011.09.006. PMID: 21978911. PMCID: PMC3210330.
Article8. Muñoz J, Stange DE, Schepers AG, van de Wetering M, Koo BK, Itzkovitz S, Volckmann R, Kung KS, Koster J, Radulescu S, Myant K, Versteeg R, Sansom OJ, van Es JH, Barker N, van Oudenaarden A, Mohammed S, Heck AJ, Clevers H. 2012; The Lgr5 intestinal stem cell signature: robust expression of proposed quiescent '+4' cell markers. EMBO J. 31:3079–3091. DOI: 10.1038/emboj.2012.166. PMID: 22692129. PMCID: PMC3400017.
Article9. Ghaleb AM, Yang VW. 2017; Krüppel-like factor 4 (KLF4): what we currently know. Gene. 611:27–37. DOI: 10.1016/j.gene.2017.02.025. PMID: 28237823. PMCID: PMC5391259.
Article10. Kuruvilla JG, Kim CK, Ghaleb AM, Bialkowska AB, Kuo CJ, Yang VW. 2016; Krüppel-like factor 4 modulates development of BMI1(+) intestinal stem cell-derived lineage following γ-radiation-induced gut injury in mice. Stem Cell Reports. 6:815–824. DOI: 10.1016/j.stemcr.2016.04.014. PMID: 27237377. PMCID: PMC4911500.
Article11. Ghaleb AM, Aggarwal G, Bialkowska AB, Nandan MO, Yang VW. 2008; Notch inhibits expression of the Krüppel-like factor 4 tumor suppressor in the intestinal epithelium. Mol Cancer Res. 6:1920–1927. DOI: 10.1158/1541-7786.MCR-08-0224. PMID: 19074836. PMCID: PMC2628949.
Article12. Ghaleb AM, McConnell BB, Kaestner KH, Yang VW. 2011; Altered intestinal epithelial homeostasis in mice with intestine-specific deletion of the Krüppel-like factor 4 gene. Dev Biol. 349:310–320. DOI: 10.1016/j.ydbio.2010.11.001. PMID: 21070761. PMCID: PMC3022386.
Article13. Imajo M, Ebisuya M, Nishida E. 2015; Dual role of YAP and TAZ in renewal of the intestinal epithelium. Nat Cell Biol. 17:7–19. DOI: 10.1038/ncb3084. PMID: 25531778.
Article14. Kim CK, Saxena M, Maharjan K, Song JJ, Shroyer KR, Bialkowska AB, Shivdasani RA, Yang VW. 2020; Krüppel-like factor 5 regulates stemness, lineage specification, and regeneration of intestinal epithelial stem cells. Cell Mol Gastroenterol Hepatol. 9:587–609. DOI: 10.1016/j.jcmgh.2019.11.009. PMID: 31778829. PMCID: PMC7078555.
Article15. Miyoshi H, Stappenbeck TS. 2013; In vitro expansion and genetic modification of gastrointestinal stem cells in spheroid cul-ture. Nat Protoc. 8:2471–2482. DOI: 10.1038/nprot.2013.153. PMID: 24232249. PMCID: PMC3969856.
Article16. Flandez M, Guilmeau S, Blache P, Augenlicht LH. 2008; KLF4 regulation in intestinal epithelial cell maturation. Exp Cell Res. 314:3712–3723. DOI: 10.1016/j.yexcr.2008.10.004. PMID: 18977346. PMCID: PMC2652355.
Article17. Itzkovitz S, Lyubimova A, Blat IC, Maynard M, van Es J, Lees J, Jacks T, Clevers H, van Oudenaarden A. 2011; Single-molecule transcript counting of stem-cell markers in the mouse intestine. Nat Cell Biol. 14:106–114. DOI: 10.1038/ncb2384. PMID: 22119784. PMCID: PMC3292866.
Article18. Yan KS, Gevaert O, Zheng GXY, Anchang B, Probert CS, Larkin KA, Davies PS, Cheng ZF, Kaddis JS, Han A, Roelf K, Calderon RI, Cynn E, Hu X, Mandleywala K, Wilhelmy J, Grimes SM, Corney DC, Boutet SC, Terry JM, Belgrader P, Ziraldo SB, Mikkelsen TS, Wang F, von Furstenberg RJ, Smith NR, Chandrakesan P, May R, Chrissy MAS, Jain R, Cartwright CA, Niland JC, Hong YK, Carrington J, Breault DT, Epstein J, Houchen CW, Lynch JP, Martin MG, Plevritis SK, Curtis C, Ji HP, Li L, Henning SJ, Wong MH, Kuo CJ. 2017; Intestinal enteroendocrine lineage cells possess homeostatic and injury-inducible stem cell activity. Cell Stem Cell. 21:78–90.e6. DOI: 10.1016/j.stem.2017.06.014. PMID: 28686870. PMCID: PMC5642297.
Article19. Schneider H, Pelaseyed T, Svensson F, Johansson MEV. 2018; Study of mucin turnover in the small intestine by in vivo labeling. Sci Rep. 8:5760. DOI: 10.1038/s41598-018-24148-x. PMID: 29636525. PMCID: PMC5893601.
Article20. Gersemann M, Becker S, Kübler I, Koslowski M, Wang G, Herrlinger KR, Griger J, Fritz P, Fellermann K, Schwab M, Wehkamp J, Stange EF. 2009; Differences in goblet cell differentiation between Crohn's disease and ulcerative colitis. Differentiation. 77:84–94. DOI: 10.1016/j.diff.2008.09.008. PMID: 19281767.
Article21. Katz JP, Perreault N, Goldstein BG, Lee CS, Labosky PA, Yang VW, Kaestner KH. 2002; The zinc-finger transcription factor Klf4 is required for terminal differentiation of goblet cells in the colon. Development. 129:2619–2628. PMID: 12015290. PMCID: PMC2225535.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Effect of BMI1 Knockdown on Cell Proliferation, Apoptosis, Invasiveness, and Migration of U251 Glioma Cells
- Proteomics analysis of tumor-infiltrated T cell reveals CD127+ and KLRG1+ memory CD8+ T cells control immunotherapy efficacy in hepatocellular carcinoma
- Human Placenta-Derived ECM Supports Tri-Lineage Differentiation of Human Induced Pluripotent Stem Cells
- Stem Cells in Colorectal Cancer: New Potential Therapeutic Target
- Biopsy Findings of Conjunotival Goblet Cell Densities Around The Pterygium