Ann Lab Med.
2020 Jul;40(4):345-347. 10.3343/alm.2020.40.4.345.
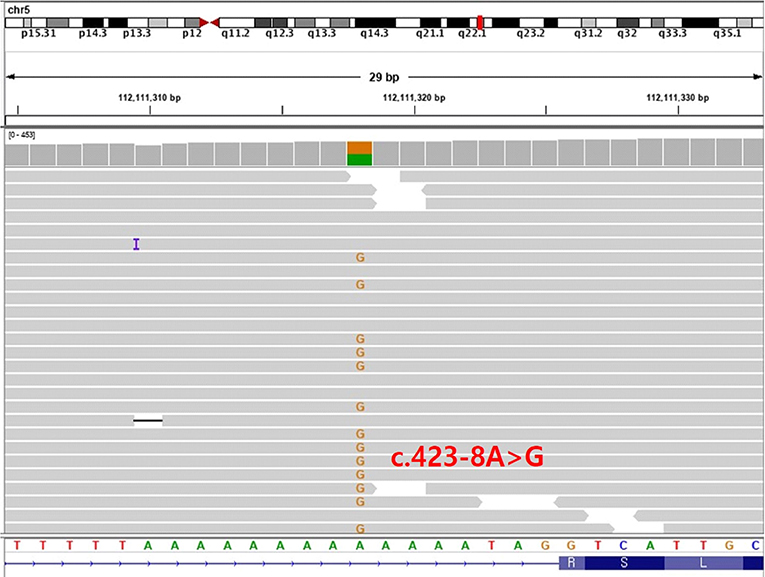
Identification of a Novel Splice Variant (c.423-8A>G) of APC by RNA Sequencing
- Affiliations
-
- 1Department of Laboratory Medicine, University of Ulsan College of Medicine and Asan Medical Center, Seoul, Korea. wlee1@amc.seoul.kr
- 2Department of Laboratory Medicine, Kangwon National University Hospital, Chuncheon, Korea.
- 3Department of Oncology, University of Ulsan College of Medicine and Asan Medical Center, Seoul, Korea.
- 4Department of Surgery, University of Ulsan College of Medicine and Asan Medical Center, Seoul, Korea.
- 5Department of Gastroenterology, University of Ulsan College of Medicine and Asan Medical Center, Seoul, Korea.
- KMID: 2470335
- DOI: http://doi.org/10.3343/alm.2020.40.4.345
Abstract
- No abstract available.
MeSH Terms
Figure
Reference
-
1. Zhang Y, Lu G, Hu Q, Wang X, Li C, Mao Y, et al. A de novo germline mutation of APC for inheritable colon cancer in a Chinese family using multigene next generation sequencing. Biochem Biophys Res Commun. 2014; 447:503–507.2. Stenson PD, Ball EV, Mort M, Phillips AD, Shiel JA, Thomas NS, et al. Human Gene Mutation Database (HGMD): 2003 update. Hum Mutat. 2003; 21:577–581.
Article3. Jian X, Boerwinkle E, Liu X. In silico prediction of splice-altering single nucleotide variants in the human genome. Nucleic Acids Res. 2014; 42:13534–13544.
Article4. Desmet FO, Hamroun D, Lalande M, Collod-Beroud G, Claustres M, Beroud C. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009; 37:e67.
Article5. Yeo G, Burge CB. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Comput Biol. 2004; 11:377–394.
Article6. Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015; 17:405–424.
Article7. Tsukanov AS, Pospekhova NI, Shubin VP, Kuzminov AM, Kashnikov VN, Frolov SA, et al. Mutations in the APC gene in Russian patients with classic form of familial adenomatous polyposis. Russ J Genet. 2017; 53:369–375.8. Jarry J, Brunet JS, Laframboise R, Drouin R, Latreille J, Richard C, et al. A survey of APC mutations in Quebec. Fam Cancer. 2011; 10:659–665.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- A Novel Splice Variant (c.438T>A) of APC, Suspected by Family History and Confirmed by RNA Sequencing
- Identification of Alternative Splicing and Fusion Transcripts in Non-Small Cell Lung Cancer by RNA Sequencing
- Functional involvement of src and focal adhesion kinase in a CD99 splice variant-induced motility of human breast cancer cells.
- A Novel Adenomatous Polyposis Coli Gene Mutation of Cribriform-Morular Variant Papillary Thyroid Carcinoma: A Case Report
- Analysis of Whole Transcriptome Sequencing Data: Workflow and Software