Lab Med Online.
2019 Apr;9(2):45-56. 10.3343/lmo.2019.9.2.45.
Effects of Pre-analytical Variables on Cell-free DNA Extraction for Liquid Biopsy
- Affiliations
-
- 1Department of Laboratory Medicine, College of Medicine, Ewha Womans University, Seoul, Korea. tdjeong@ewha.ac.kr
- 2Division of Pulmonary and Critical Care Medicine, Department of Internal Medicine, College of Medicine, Ewha Womans University, Seoul, Korea.
- KMID: 2441823
- DOI: http://doi.org/10.3343/lmo.2019.9.2.45
Abstract
- BACKGROUND
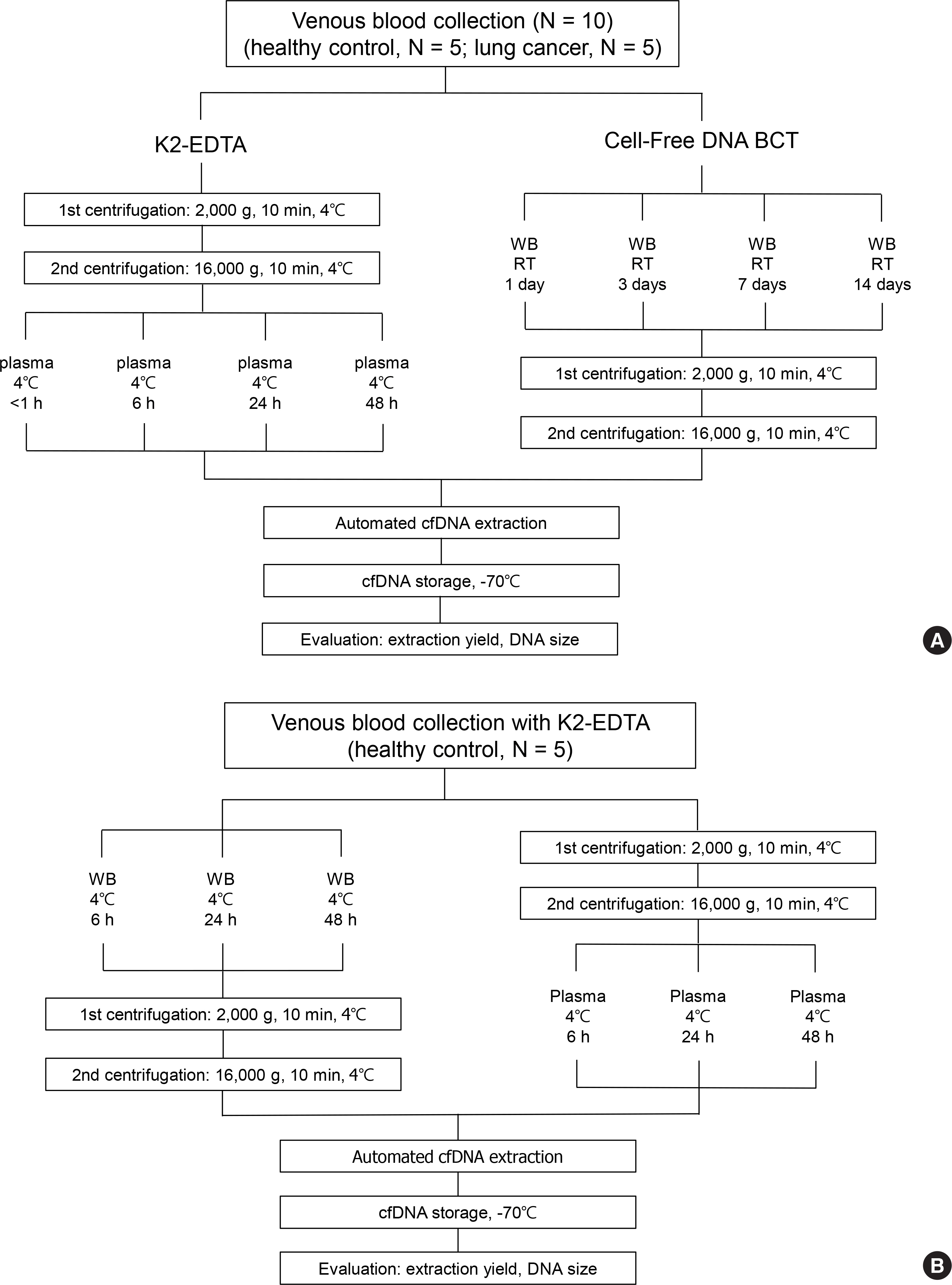
Extraction of cell-free DNA (cfDNA) is a key step for determining the quality of cfDNA-related molecular diagnostics. We evaluated the effect of sample containers and sample storage conditions on cfDNA extraction.
METHODS
The cfDNA extraction using the MagMAX Cell-Free DNA Isolation Kit from five healthy controls and five lung cancer patients was evaluated according to the type of sample container and storage conditions: K2-EDTA container, <1, 6, 24, and 48 hr storage at 4℃ after immediate plasma separation; and Cell-Free DNA BCT container, <1, 3, 7, and 14 days stored at room temperature. Mutation analysis of EGFR exons 18-21 was performed. To assess the effect of a delay in centrifugation, EDTA whole blood samples from five healthy individuals were stored at 4℃ for 6, 12, and 24 hr before plasma separation.
RESULTS
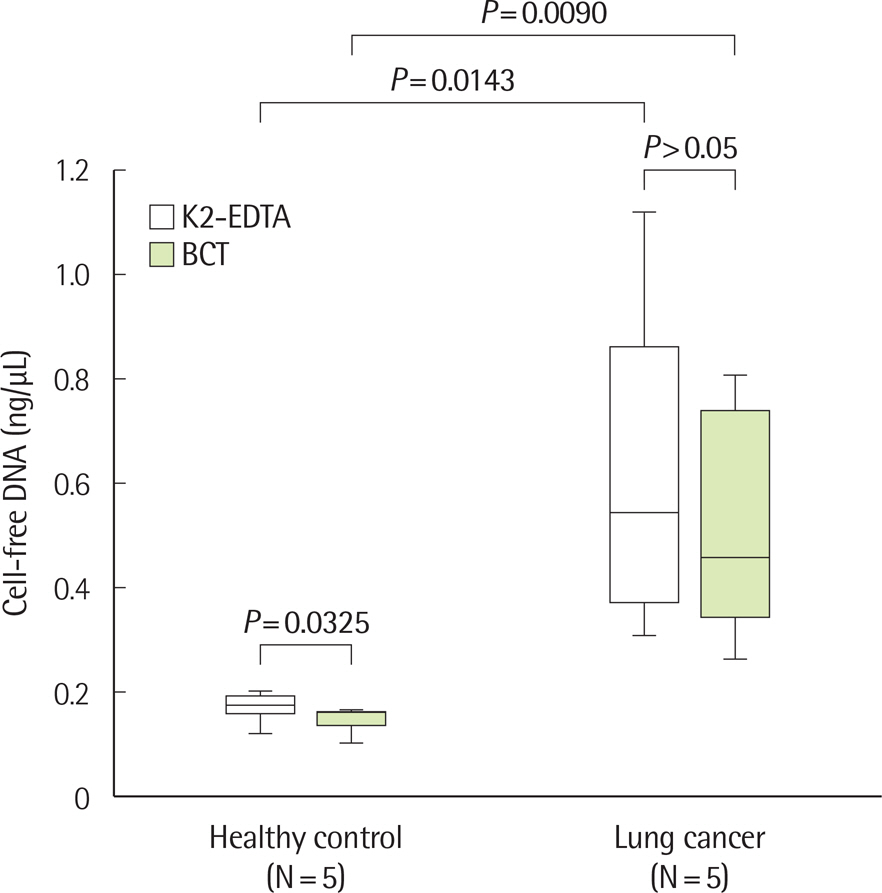
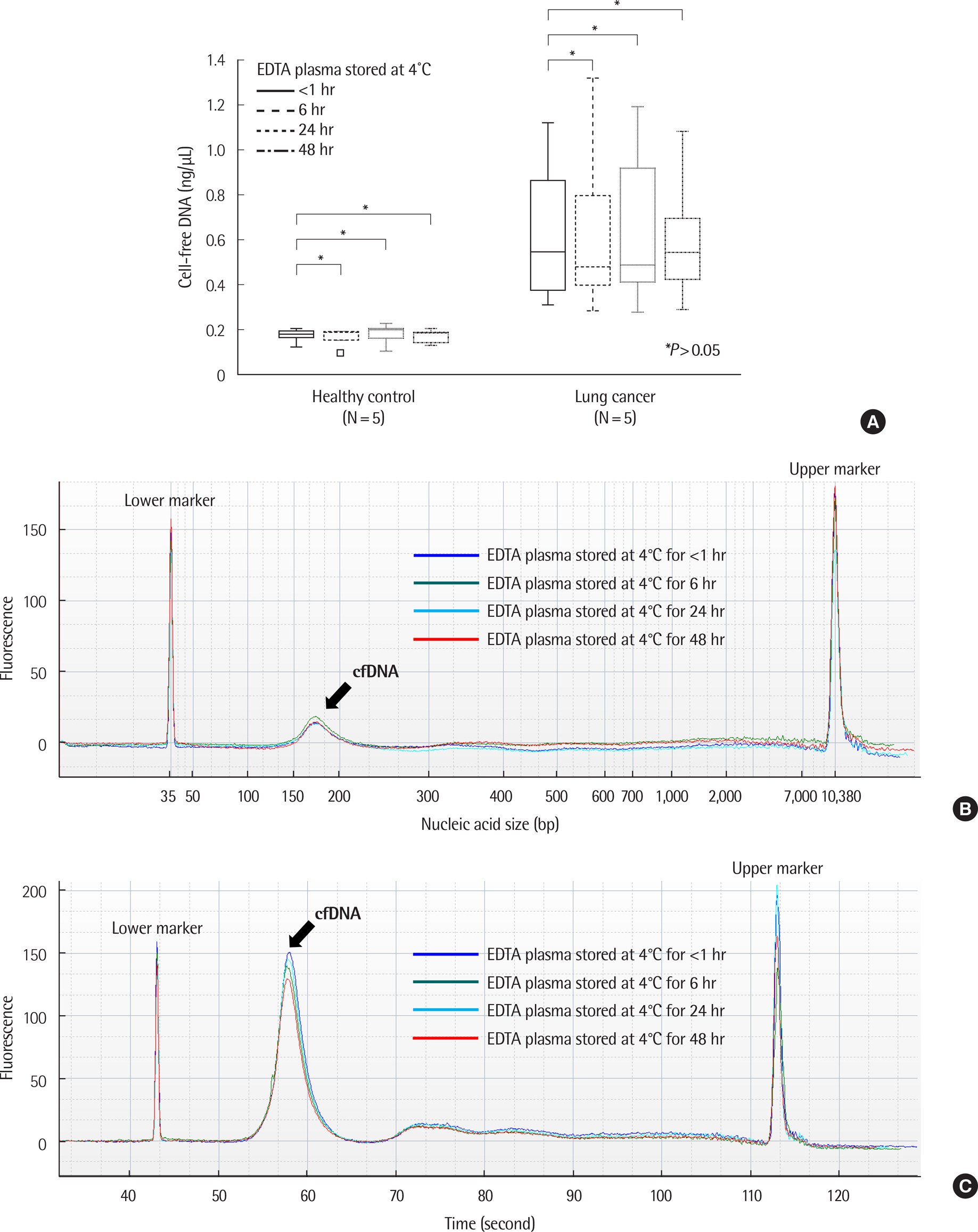
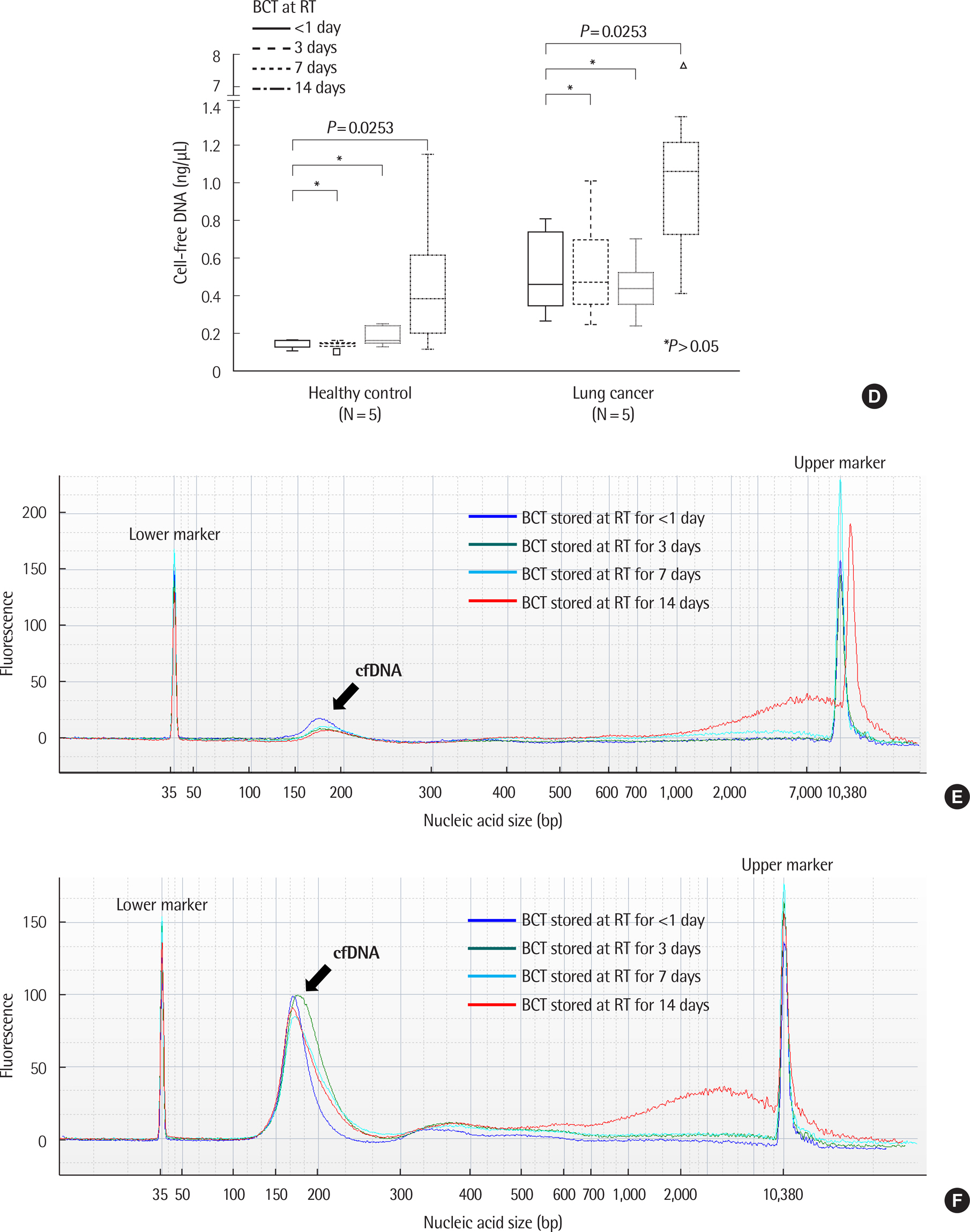
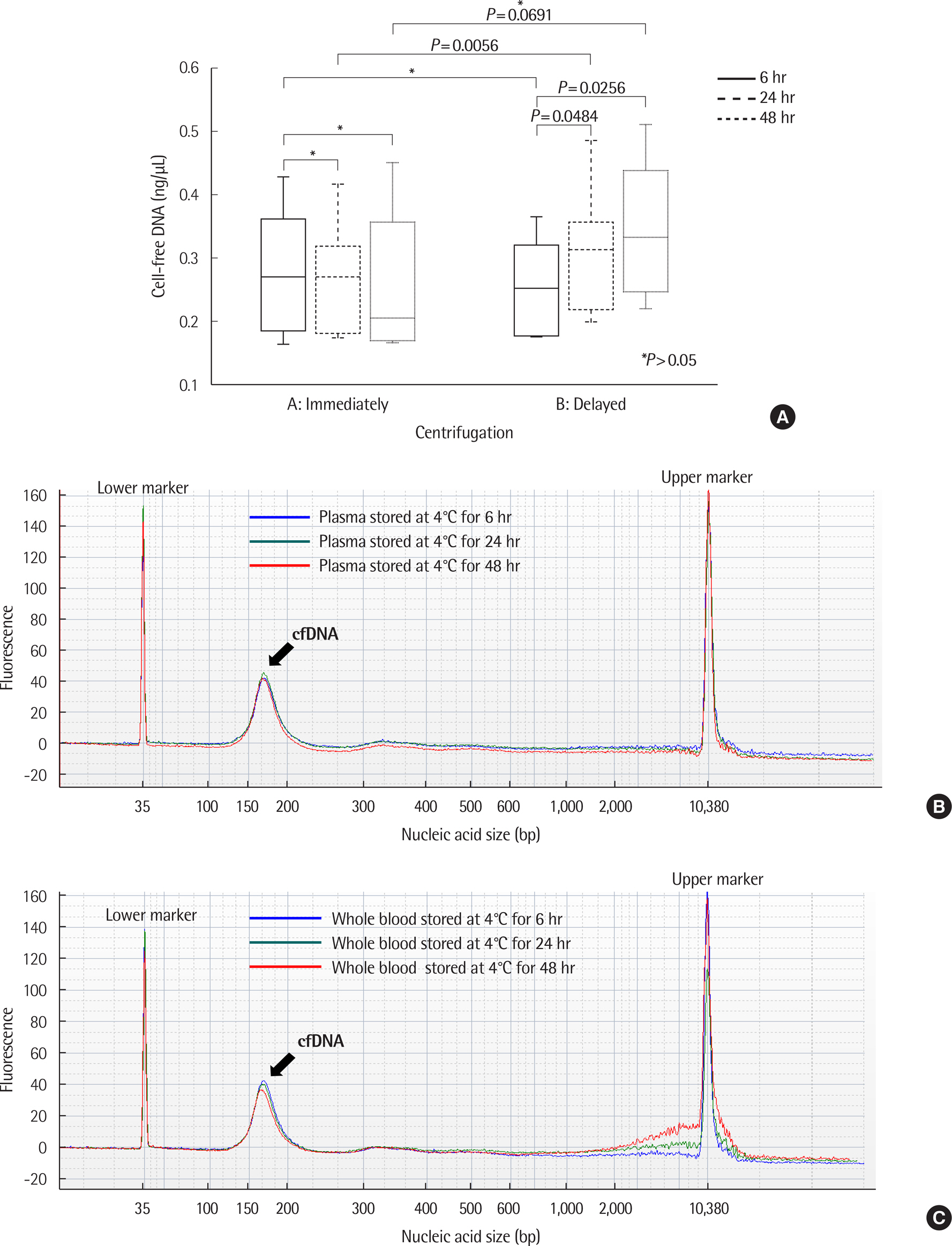
There was no significant difference in the amount and nucleic acid size of cfDNA in both controls and patients with cancer when EDTA plasma was stored at 4℃ up to 48 hr. The amount and size of cfDNA in the BCT container were not different up to 7 days; however, the 14-day sample showed an increase in cfDNA concentration due to genomic DNA contamination. EGFR mutations were detected on EDTA containers up to 48 hr and with BCT containers up to 14 days. When EDTA whole blood was stored at 4℃ and plasma separation was delayed, the cfDNA concentration increased from 24 hr.
CONCLUSIONS
The cfDNA extraction was affected by the sample containers and storage conditions.
Keyword
MeSH Terms
Figure
Cited by 2 articles
-
Targeted Next-Generation Sequencing of Plasma Cell-Free DNA in Korean Patients with Hepatocellular Carcinoma
Hyojin Chae, Pil Soo Sung, Hayoung Choi, Ahlm Kwon, Dain Kang, Yonggoo Kim, Myungshin Kim, Seung Kew Yoon
Ann Lab Med. 2021;41(2):198-206. doi: 10.3343/alm.2021.41.2.198.Clinical Practice Guidelines for Pre-Analytical Procedures of Plasma Epidermal Growth Factor Receptor Variant Testing
Saeam Shin, Hye In Woo, Jong-Won Kim, Yoonjung Kim, Kyung-A Lee
Ann Lab Med. 2022;42(2):141-149. doi: 10.3343/alm.2022.42.2.141.
Reference
-
References
1. Volik S, Alcaide M, Morin RD, Collins C. Cell-free DNA (cfDNA): Clinical signifcance and utility in cancer shaped by emerging technologies. Mol Cancer Res. 2016; 14:898–908.2. Wan JCM, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Cal-das C, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat Rev Cancer. 2017; 17:223–38.
Article3. Maheswaran S, Sequist LV, Nagrath S, Ulkus L, Brannigan B, Collura CV, et al. Detection of mutations in EGFR in circulating lung-cancer cells. N Engl J Med. 2008; 359:366–77.4. Kwapisz D. The frst liquid biopsy test approved. Is it a new era of mutation testing for non-small cell lung cancer? Ann Transl Med. 2017; 5:46.5. Diaz LA Jr and Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014; 32:579–86.6. Ilie M and Hofman P. Pros: Can tissue biopsy be replaced by liquid biopsy? Transl Lung Cancer Res. 2016; 5:420–3.7. Krishnamurthy N, Spencer E, Torkamani A, Nicholson L. Liquid biopsies for cancer: Coming to a patient near you. J Clin Med. 2017; 6:3.
Article8. Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J, Endesfelder D, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012; 366:883–92.
Article9. Bedard PL, Hansen AR, Ratain MJ, Siu LL. Tumour heterogeneity in the clinic. Nature. 2013; 501:355–64.
Article10. Thierry AR, El Messaoudi S, Gahan PB, Anker P, Stroun M. Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev. 2016; 35:347–76.
Article11. Yao W, Mei C, Nan X, Hui L. Evaluation and comparison of in vitro degradation kinetics of DNA in serum, urine and saliva: A qualitative study. Gene. 2016; 590:142–8.
Article12. Bronkhorst AJ, Aucamp J, Pretorius PJ. Cell-free DNA: Preanalytical variables. Clin Chim Acta. 2015; 450:243–53.
Article13. Heitzer E, Ulz P, Geigl JB. Circulating tumor DNA as a liquid biopsy for cancer. Clin Chem. 2015; 61:112–23.
Article14. Chen W, Cai F, Zhang B, Zhong XY. Strategies of reducing input sample volume for extracting circulating cell-free nuclear DNA and mitochondrial DNA in plasma. Clin Chem Lab Med. 2011; 50:261–5.
Article15. Devonshire AS, Whale AS, Gutteridge A, Jones G, Cowen S, Foy CA, et al. Towards standardisation of cell-free DNA measurement in plasma: controls for extraction effciency, fragment size bias and quantifcation. Anal Bioanal Chem. 2014; 406:6499–512.16. van Dessel LF, Beije N, Helmijr JC, Vitale SR, Kraan J, Look MP, et al. Application of circulating tumor DNA in prospective clinical oncology trials – standardization of preanalytical conditions. Mol Oncol. 2017; 11:295–304.
Article17. Medina Diaz I, Nocon A, Mehnert DH, Fredebohm J, Diehl F, Holtrup F. Performance of Streck cfDNA blood collection tubes for liquid biopsy testing. PLoS One. 2016; 11:e0166354.
Article18. Sherwood JL, Corcoran C, Brown H, Sharpe AD, Musilova M, Kohl-mann A. Optimised preanalytical methods improve KRAS mutation detection in circulating tumour DNA (ctDNA) from patients with nonsmall cell lung cancer (NSCLC). PLoS One. 2016; 11:e0150197.
Article19. Parpart-Li S, Bartlett B, Popoli M, Adleff V, Tucker L, Steinberg R, et al. The effect of preservative and temperature on the analysis of circulating tumor DNA. Clin Cancer Res. 2017; 23:2471–7.
Article20. van Ginkel JH, van den Broek DA, van Kuik J, Linders D, de Weger R, Willems SM, et al. Preanalytical blood sample workup for cell-free DNA analysis using Droplet Digital PCR for future molecular cancer diagnostics. Cancer Med. 2017; 6:2297–307.
Article21. Warton K, Yuwono NL, Cowley MJ, McCabe MJ, So A, Ford CE. Evaluation of Streck BCT and PAXgene stabilised blood collection tubes for cell-free circulating DNA studies in plasma. Mol Diagn Ther. 2017; 21:563–70.
Article22. Streck. Cell-Free DNA BCT: INSTRUCTIONS FOR USE. Omaha N, USA;. 2014. https://www.streck.com/collection/cell-free-dna-bct/. (Assessed on Jan 2018).23. Alix-Panabieres C and Pantel K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016; 6:479–91.
Article24. Zhang R, Peng R, Li Z, Gao P, Jia S, Yang X, et al. Synthetic circulating cell-free DNA as quality control materials for somatic mutation detection in liquid biopsy for cancer. Clin Chem. 2017; 63:1465–75.
Article25. Xue X, Teare MD, Holen I, Zhu YM, Woll PJ. Optimizing the yield and utility of circulating cell-free DNA from plasma and serum. Clin Chim Acta. 2009; 404:100–4.
Article26. Leung F, Kulasingam V, Diamandis EP, Hoon DS, Kinzler K, Pantel K, et al. Circulating tumor DNA as a cancer biomarker: fact or fction? Clin Chem. 2016; 62:1054–60.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Clinical Application of Circulating Tumor DNA Analysis
- Liquid biopsy for early detection and therapeutic monitoring of hepatocellular carcinoma
- Liquid biopsy? A recent breakthrough in noninvasive bladder cancer surveillance
- Role of Liquid Biopsies in Colorectal Cancer
- Cell-Free DNA in Oncology: Gearing up for Clinic