Yonsei Med J.
2018 Jul;59(5):652-661. 10.3349/ymj.2018.59.5.652.
A New Integrated Newborn Screening Workflow Can Provide a Shortcut to Differential Diagnosis and Confirmation of Inherited Metabolic Diseases
- Affiliations
-
- 1Department of Pediatrics, Seoul National University Children's Hospital, Seoul National University College of Medicine, Seoul, Korea.
- 2SD Genomics Co., Ltd., Seoul, Korea.
- 3Department of Laboratory Medicine, Yonsei University College of Medicine, Seoul, Korea. KAL1119@yuhs.ac
- 4Department of Pediatrics, Yonsei University College of Medicine, Seoul, Korea.
- 5Department of Laboratory Medicine and Genetics, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea. kimjw@skku.edu
- KMID: 2412656
- DOI: http://doi.org/10.3349/ymj.2018.59.5.652
Abstract
- PURPOSE
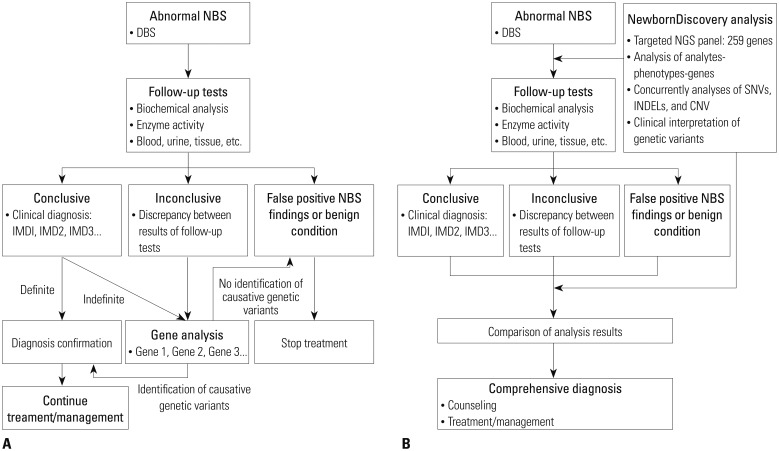
We developed a new workflow design which included results from both biochemical and targeted gene sequencing analysis interpreted comprehensively. We then conducted a pilot study to evaluate the benefit of this new approach in newborn screening (NBS) and demonstrated the efficiency of this workflow in detecting causative genetic variants.
MATERIALS AND METHODS
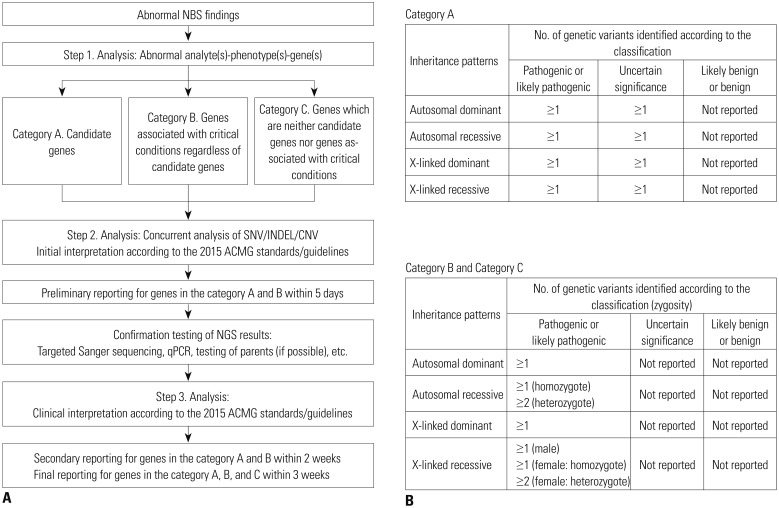
Ten patients in Group 1 were diagnosed clinically using biochemical assays only, and 10 newborns in Group 2 were diagnosed with suspected inherited metabolic disease (IMD) in NBS. We applied NewbornDiscovery (SD Genomics), an integrated workflow design that encompasses analyte-phenotype-gene, single nucleotide variant/small insertion and deletion/copy number variation analyses along with clinical interpretation of genetic variants related to each participant's condition.
RESULTS
A molecular genetic diagnosis was established in 95% (19/20) of individuals. In Group 1, 13 and 7 of 20 alleles were classified as pathogenic and likely pathogenic, respectively. In Group 2, 11 and 6 of 17 alleles with identified causative variants were pathogenic and likely pathogenic, respectively. There were no variants of uncertain significance. For each individual, the NewbornDiscovery and biochemical analysis results reached 100% concordance, since the single newborn testing negative for causative genetic variant in Group 2 showed a benign clinical course.
CONCLUSION
This integrated diagnostic workflow resulted in a high yield. This approach not only enabled early confirmation of specific IMD, but also detected conditions not included in the current NBS.
Keyword
MeSH Terms
Figure
Reference
-
1. Park KJ, Kim JW. Perspectives on next-generation newborn screening. Lab Med Online. 2015; 5:169–175.
Article2. Mak CM, Lee HC, Chan AY, Lam CW. Inborn errors of metabolism and expanded newborn screening: review and update. Crit Rev Clin Lab Sci. 2013; 50:142–162. PMID: 24295058.
Article3. Schulze A, Lindner M, Kohlmüller D, Olgemöller K, Mayatepek E, Hoffmann GF. Expanded newborn screening for inborn errors of metabolism by electrospray ionization-tandem mass spectrometry: results, outcome, and implications. Pediatrics . 2003; 111(6 Pt 1):1399–1406. PMID: 12777559.
Article4. Bhattacharjee A, Sokolsky T, Wyman SK, Reese MG, Puffenberger E, Strauss K, et al. Development of DNA confirmatory and high-risk diagnostic testing for newborns using targeted next-generation DNA sequencing. Genet Med. 2015; 17:337–347. PMID: 25255367.
Article5. Bodian DL, Klein E, Iyer RK, Wong WS, Kothiyal P, Stauffer D, et al. Utility of whole-genome sequencing for detection of newborn screening disorders in a population cohort of 1,696 neonates. Genet Med. 2016; 18:221–230. PMID: 26334177.
Article6. Hollegaard MV, Grauholm J, Nielsen R, Grove J, Mandrup S, Hougaard DM. Archived neonatal dried blood spot samples can be used for accurate whole genome and exome-targeted next-generation sequencing. Mol Genet Metab. 2013; 110:65–72. PMID: 23830478.
Article7. Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015; 17:405–424. PMID: 25741868.
Article8. Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010; 26:589–595. PMID: 20080505.
Article9. McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010; 20:1297–1303. PMID: 20644199.
Article10. Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010; 38:e164. PMID: 20601685.
Article11. Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009; 4:1073–1081. PMID: 19561590.
Article12. Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010; 7:248–249. PMID: 20354512.
Article13. Schwarz JM, Cooper DN, Schuelke M, Seelow D. Mutation-Taster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014; 11:361–362. PMID: 24681721.
Article14. Shihab HA, Gough J, Cooper DN, Stenson PD, Barker GL, Edwards KJ, et al. Predicting the functional, molecular, and phenotypic consequences of amino acid substitutions using hidden Markov models. Hum Mutat. 2013; 34:57–65. PMID: 23033316.
Article15. Jian X, Boerwinkle E, Liu X. In silico prediction of splice-altering single nucleotide variants in the human genome. Nucleic Acids Res. 2014; 42:13534–13544. PMID: 25416802.
Article16. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001; 25:402–408. PMID: 11846609.17. Kim YM, Kim JH, Choi JH, Kim GH, Kim JM, Kang M, et al. Determination of autosomal dominant or recessive methionine adenosyltransferase I/III deficiencies based on clinical and molecular studies. Mol Med. 2016; 22:147–155.
Article18. Chien YH, Abdenur JE, Baronio F, Bannick AA, Corrales F, Couce M, et al. Mudd's disease (MAT I/III deficiency): a survey of data for MAT1A homozygotes and compound heterozygotes. Orphanet J Rare Dis. 2015; 10:99. PMID: 26289392.
Article19. Fernández-Irigoyen J, Santamaría E, Chien YH, Hwu WL, Korman SH, Faghfoury H, et al. Enzymatic activity of methionine adenosyltransferase variants identified in patients with persistent hypermethioninemia. Mol Genet Metab. 2010; 101:172–177. PMID: 20675163.
Article20. Lee YW, Lee DH, Vockley J, Kim ND, Lee YK, Ki CS. Different spectrum of mutations of isovaleryl-CoA dehydrogenase (IVD) gene in Korean patients with isovaleric acidemia. Mol Genet Metab. 2007; 92:71–77. PMID: 17576084.
Article21. Kim YM, Cheon CK, Park KH, Park S, Kim GH, Yoo HW, et al. Novel and recurrent ACADS mutations and clinical manifestations observed in Korean patients with short-chain acyl-coenzyme a dehydrogenase deficiency. Ann Clin Lab Sci. 2016; 46:360–366. PMID: 27466294.22. Shirao K, Okada S, Tajima G, Tsumura M, Hara K, Yasunaga S, et al. Molecular pathogenesis of a novel mutation, G108D, in short-chain acyl-CoA dehydrogenase identified in subjects with short-chain acyl-CoA dehydrogenase deficiency. Hum Genet. 2010; 127:619–628. PMID: 20376488.
Article23. Magoulas PL, El-Hattab AW. Systemic primary carnitine deficiency: an overview of clinical manifestations, diagnosis, and management. Orphanet J Rare Dis. 2012; 7:68. PMID: 22989098.
Article24. Bennett MJ, Santani AB. Carnitine palmitoyltransferase 1A deficiency. In : Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, editors. GeneReviews(R). Seattle (WA): University of Washington;1993.25. Goodman SI, Binard RJ, Woontner MR, Frerman FE. Glutaric acidemia type II: gene structure and mutations of the electron transfer flavoprotein:ubiquinone oxidoreductase (ETF:QO) gene. Mol Genet Metab. 2002; 77:86–90. PMID: 12359134.
Article26. Filippo CA, Ardon O, Longo N. Glycosylation of the OCTN2 carnitine transporter: study of natural mutations identified in patients with primary carnitine deficiency. Biochim Biophys Acta. 2011; 1812:312–320. PMID: 21126579.
Article27. Wiltink RC, Kruijshaar ME, van Minkelen R, Onkenhout W, Verheijen FW, Kemper EA, et al. Neonatal screening for profound biotinidase deficiency in the Netherlands: consequences and considerations. Eur J Hum Genet. 2016; 24:1424–1429. PMID: 27329734.
Article28. Therrell BL, Padilla CD, Loeber JG, Kneisser I, Saadallah A, Borrajo GJ, et al. Current status of newborn screening worldwide: 2015. Semin Perinatol. 2015; 39:171–187. PMID: 25979780.
Article29. Howard HC, Knoppers BM, Cornel MC, Wright Clayton E, Sénécal K, Borry P, et al. Whole-genome sequencing in newborn screening? A statement on the continued importance of targeted approaches in newborn screening programmes. Eur J Hum Genet. 2015; 23:1593–1600. PMID: 25626707.
Article30. Berg JS, Powell CM. Potential uses and inherent challenges of using genome-scale sequencing to augment current newborn screening. Cold Spring Harb Perspect Med. 2015; 5:a023150. PMID: 26438605.
Article31. Kim SH, Park HD, Sohn YB, Park SW, Cho SY, Ji S, et al. Mutations of ACADS gene associated with short-chain acyl-coenzyme A dehydrogenase deficiency. Ann Clin Lab Sci. 2011; 41:84–88. PMID: 21325261.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Screening newborns for metabolic disorders based on targeted metabolomics using tandem mass spectrometry
- The prevalence of pediatric endocrine and metabolic diseases in Korea
- Seven-year experience with inherited metabolic disorders screening by tandem mass spectrometry
- A Population-Based Genomic Study of Inherited Metabolic Diseases Detected Through Newborn Screening
- Perspectives on Next-Generation Newborn Screening